| 出品公司: | Clontech |
|---|---|
| 载体名称: | pIRES2-EGFP, pIRES2 EGFP |
| 质粒类型: | 哺乳动物表达载体;双顺反子载体;荧光报告载体 |
| 启动子: | CMV |
| 表达水平: | 高 |
| 克隆方法: | 多克隆位点,限制性内切酶 |
| 载体大小: | 5308 bp |
| 5' 测序引物: | CMV-F: 5'-CGCAAATGGGCGGTAGGCGTG-3' |
| 3' 测序引物: | EGFP-N: 5'-CGTCGCCGTCCAGCTCGACCAG-3' |
| 载体标签: | 绿色荧光蛋白EGFP |
| 载体抗性: | 卡那 |
| 筛选标记: | 新霉素(G418) |
| 备注: |
通过一个转录mRNA,能够同时表达EGFP蛋白和一个目的基因。
|
| 产品目录号: | 6029-1 |
| 稳定性: | 稳表达 或 瞬表达 |
| 组成型: | 组成型 |
| 病毒/非病毒: | 非病毒 |
买家导航

pIRES2-EGFP载体 pIRES2 EGFP质粒基本信息
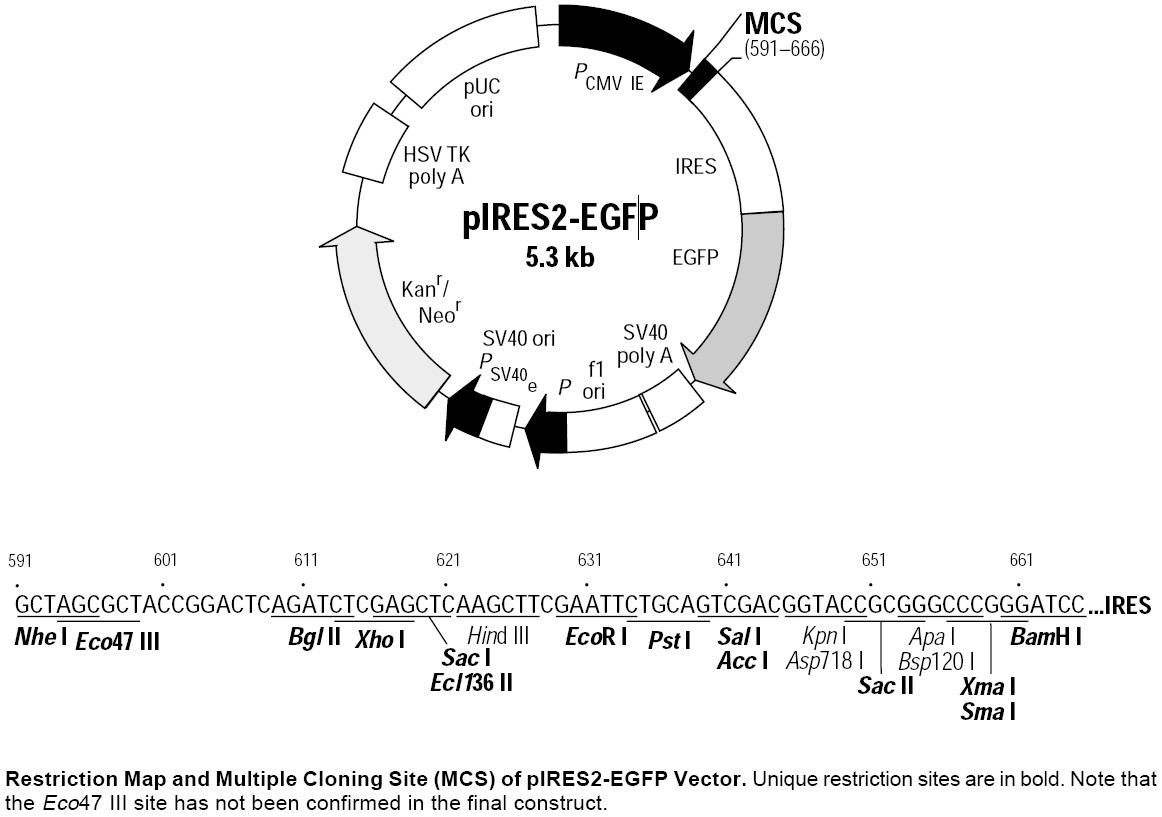
pIRES2-EGFP质粒图谱 pIRES2 EGFP载体图谱和pIRES2-EGFP载体序列 pIRES2 EGFP质粒序列多克隆位点信息
pIRES2-EGFP质粒 pIRES2 EGFP载体简介
Description:
pIRES2-EGFP contains the internal ribosome entry site (IRES; 1, 2) of the encephalomyocarditis virus (ECMV) between the MCS and the enhanced green fluorescent protein (EGFP) coding region. This permits both the gene of interest (cloned into the MCS) and the EGFP gene to be translated from a single bicistronic mRNA. pIRES2-EGFP is designed for the efficient selection (by flow cytometry or other methods) of transiently transfected mammalian cells expressing EGFP and the protein of interest. This vector can also be used to express EGFP alone or to obtain stably transfected cell lines without time-consuming drug and clonal selection. EGFP is a red-shifted variant of wild-type GFP (3–5) which has been optimized for brighter fluorescence and higher expression in mammalian cells. (Excitation maximum = 488 nm; emission maximum = 507 nm.) EGFP encodes the GFPmut1 variant (6) which contains the double-amino-acid substitution of Phe-64 to Leu and Ser-65 to Thr. The coding sequence of the EGFP gene contains more than 190 silent base changes which correspond to human codon-usage preferences (7). Sequences flanking EGFP have been converted to a Kozak consensus translation initiation site (8) to further increase the translation efficiency in eukaryotic cells. The MCS in pIRES2-EGFP is between the immediate early promoter of cytomegalovirus (PCMV IE) and the IRES sequence. SV40 polyadenylation signals downstream of the EGFP gene direct proper processing of the 3' end of the bicistronic mRNA. The vector backbone also contains an SV40 origin for replication in mammalian cells expressing the SV40 T antigen. A neomycin-resistance cassette (Neor), consisting of the SV40 early promoter, the neomycin/kanamycin resistance gene of Tn5, and polyadenylation signals from the herpes simplex virus thymidine kinase (HSV TK) gene, allows stably transfected eukaryotic cells to be selected using G418. A bacterial promoter upstream of this cassette expresses kanamycin resistance in E. coli. The pIRES2-EGFP backbone also provides a pUC origin of replication for propagation in E. coli and an f1 origin for single-stranded DNA production. pIRES2-EGFP replaces (but is not derived from) the pIRES-EGFP Vector previously sold by BD Biosciences Clontech. pIRES2-EGFP is functionally similarly to pIRES-EGFP; however, pIRES2- EGFP gives brighter EGFP fluorescence than the older vector. Note that the Xba I site at position 1987 is methylated in the DNA provided by BD Biosciences Clontech. If you wish to digest the vector with this enzyme, you will need to transform the vector into a dam– host and make fresh DNA.
Use:
Genes inserted into the MCS should include the initiating ATG codon. pIRES2-EGFP and its derivatives can be introduced into mammalian cells using any standard transfection method. If required, stable transformants can be selected using G418 (9).
pIRES2-EGFP质粒序列 pIRES2 EGFP载体序列
LOCUS pIRES2-EGFP 5308 bp DNA circular SYN DEFINITION pIRES2-EGFP ACCESSION KEYWORDS SOURCE ORGANISM other sequences; artificial sequences; vectors. COMMENT This file is created by Vector NTI http://www.biofeng.com/ COMMENT ORIGDB|GenBank COMMENT VNTAUTHORNAME|biofeng.com| FEATURES Location/Qualifiers source 1..5308 /organism="pIRES2-EGFP" /mol_type="other DNA" promoter 10..562 /label="CMV_immearly_promoter" misc_feature 65..352 /label="CAG_enhancer" misc_feature 519..539 /label="CMV_fwd_primer" promoter 520..589 /label="CMV_promoter" misc_feature 666..1250 /label="IRES" gene 1242..1973 /label="EGFP" /gene="EGFP" misc_feature complement(1299..1320) /label="EGFP_N_primer" misc_feature 1437..1466 /label="Y66 (EGFP)" misc_feature 1907..1928 /label="EGFP_C_primer" misc_feature 2187..2206 /label="EBV_rev_primer" rep_origin complement(2356..2662) /label="f1_origin" promoter 2741..2769 /label="AmpR_promoter" misc_feature complement(2835..2855) /label="pBABE_3_primer" misc_feature complement(2841..3056) /label="SV40_enhancer" promoter 2853..3121 /label="SV40_promoter" rep_origin 3020..3097 /label="SV40_origin" misc_feature 3082..3101 /label="SV40pro_F_primer" CDS 3204..3998 /label="ORF frame 3" gene 3207..3995 /label="NeoR/KanR" /gene="NeoR/KanR" CDS complement(3513..4049) /label="ORF frame 3" terminator 4173..4442 /label="TK_PA_terminator" rep_origin 4590..5209 /label="pBR322_origin" ORIGIN 1 TAGTTATTAA TAGTAATCAA TTACGGGGTC ATTAGTTCAT AGCCCATATA TGGAGTTCCG 61 CGTTACATAA CTTACGGTAA ATGGCCCGCC TGGCTGACCG CCCAACGACC CCCGCCCATT 121 GACGTCAATA ATGACGTATG TTCCCATAGT AACGCCAATA GGGACTTTCC ATTGACGTCA 181 ATGGGTGGAG TATTTACGGT AAACTGCCCA CTTGGCAGTA CATCAAGTGT ATCATATGCC 241 AAGTACGCCC CCTATTGACG TCAATGACGG TAAATGGCCC GCCTGGCATT ATGCCCAGTA 301 CATGACCTTA TGGGACTTTC CTACTTGGCA GTACATCTAC GTATTAGTCA TCGCTATTAC 361 CATGGTGATG CGGTTTTGGC AGTACATCAA TGGGCGTGGA TAGCGGTTTG ACTCACGGGG 421 ATTTCCAAGT CTCCACCCCA TTGACGTCAA TGGGAGTTTG TTTTGGCACC AAAATCAACG 481 GGACTTTCCA AAATGTCGTA ACAACTCCGC CCCATTGACG CAAATGGGCG GTAGGCGTGT 541 ACGGTGGGAG GTCTATATAA GCAGAGCTGG TTTAGTGAAC CGTCAGATCC GCTAGCGCTA 601 CCGGACTCAG ATCTCGAGCT CAAGCTTCGA ATTCTGCAGT CGACGGTACC GCGGGCCCGG 661 GATCCGCCCC TCTCCCTCCC CCCCCCCTAA CGTTACTGGC CGAAGCCGCT TGGAATAAGG 721 CCGGTGTGCG TTTGTCTATA TGTTATTTTC CACCATATTG CCGTCTTTTG GCAATGTGAG 781 GGCCCGGAAA CCTGGCCCTG TCTTCTTGAC GAGCATTCCT AGGGGTCTTT CCCCTCTCGC 841 CAAAGGAATG CAAGGTCTGT TGAATGTCGT GAAGGAAGCA GTTCCTCTGG AAGCTTCTTG 901 AAGACAAACA ACGTCTGTAG CGACCCTTTG CAGGCAGCGG AACCCCCCAC CTGGCGACAG 961 GTGCCTCTGC GGCCAAAAGC CACGTGTATA AGATACACCT GCAAAGGCGG CACAACCCCA 1021 GTGCCACGTT GTGAGTTGGA TAGTTGTGGA AAGAGTCAAA TGGCTCTCCT CAAGCGTATT 1081 CAACAAGGGG CTGAAGGATG CCCAGAAGGT ACCCCATTGT ATGGGATCTG ATCTGGGGCC 1141 TCGGTGCACA TGCTTTACAT GTGTTTAGTC GAGGTTAAAA AAACGTCTAG GCCCCCCGAA 1201 CCACGGGGAC GTGGTTTTCC TTTGAAAAAC ACGATGATAA TATGGCCACA ACCATGGTGA 1261 GCAAGGGCGA GGAGCTGTTC ACCGGGGTGG TGCCCATCCT GGTCGAGCTG GACGGCGACG 1321 TAAACGGCCA CAAGTTCAGC GTGTCCGGCG AGGGCGAGGG CGATGCCACC TACGGCAAGC 1381 TGACCCTGAA GTTCATCTGC ACCACCGGCA AGCTGCCCGT GCCCTGGCCC ACCCTCGTGA 1441 CCACCCTGAC CTACGGCGTG CAGTGCTTCA GCCGCTACCC CGACCACATG AAGCAGCACG 1501 ACTTCTTCAA GTCCGCCATG CCCGAAGGCT ACGTCCAGGA GCGCACCATC TTCTTCAAGG 1561 ACGACGGCAA CTACAAGACC CGCGCCGAGG TGAAGTTCGA GGGCGACACC CTGGTGAACC 1621 GCATCGAGCT GAAGGGCATC GACTTCAAGG AGGACGGCAA CATCCTGGGG CACAAGCTGG 1681 AGTACAACTA CAACAGCCAC AACGTCTATA TCATGGCCGA CAAGCAGAAG AACGGCATCA 1741 AGGTGAACTT CAAGATCCGC CACAACATCG AGGACGGCAG CGTGCAGCTC GCCGACCACT 1801 ACCAGCAGAA CACCCCCATC GGCGACGGCC CCGTGCTGCT GCCCGACAAC CACTACCTGA 1861 GCACCCAGTC CGCCCTGAGC AAAGACCCCA ACGAGAAGCG CGATCACATG GTCCTGCTGG 1921 AGTTCGTGAC CGCCGCCGGG ATCACTCTCG GCATGGACGA GCTGTACAAG TAAAGCGGCC 1981 GCGACTCTAG ATCATAATCA GCCATACCAC ATTTGTAGAG GTTTTACTTG CTTTAAAAAA 2041 CCTCCCACAC CTCCCCCTGA ACCTGAAACA TAAAATGAAT GCAATTGTTG TTGTTAACTT 2101 GTTTATTGCA GCTTATAATG GTTACAAATA AAGCAATAGC ATCACAAATT TCACAAATAA 2161 AGCATTTTTT TCACTGCATT CTAGTTGTGG TTTGTCCAAA CTCATCAATG TATCTTAAGG 2221 CGTAAATTGT AAGCGTTAAT ATTTTGTTAA AATTCGCGTT AAATTTTTGT TAAATCAGCT 2281 CATTTTTTAA CCAATAGGCC GAAATCGGCA AAATCCCTTA TAAATCAAAA GAATAGACCG 2341 AGATAGGGTT GAGTGTTGTT CCAGTTTGGA ACAAGAGTCC ACTATTAAAG AACGTGGACT 2401 CCAACGTCAA AGGGCGAAAA ACCGTCTATC AGGGCGATGG CCCACTACGT GAACCATCAC 2461 CCTAATCAAG TTTTTTGGGG TCGAGGTGCC GTAAAGCACT AAATCGGAAC CCTAAAGGGA 2521 GCCCCCGATT TAGAGCTTGA CGGGGAAAGC CGGCGAACGT GGCGAGAAAG GAAGGGAAGA 2581 AAGCGAAAGG AGCGGGCGCT AGGGCGCTGG CAAGTGTAGC GGTCACGCTG CGCGTAACCA 2641 CCACACCCGC CGCGCTTAAT GCGCCGCTAC AGGGCGCGTC AGGTGGCACT TTTCGGGGAA 2701 ATGTGCGCGG AACCCCTATT TGTTTATTTT TCTAAATACA TTCAAATATG TATCCGCTCA 2761 TGAGACAATA ACCCTGATAA ATGCTTCAAT AATATTGAAA AAGGAAGAGT CCTGAGGCGG 2821 AAAGAACCAG CTGTGGAATG TGTGTCAGTT AGGGTGTGGA AAGTCCCCAG GCTCCCCAGC 2881 AGGCAGAAGT ATGCAAAGCA TGCATCTCAA TTAGTCAGCA ACCAGGTGTG GAAAGTCCCC 2941 AGGCTCCCCA GCAGGCAGAA GTATGCAAAG CATGCATCTC AATTAGTCAG CAACCATAGT 3001 CCCGCCCCTA ACTCCGCCCA TCCCGCCCCT AACTCCGCCC AGTTCCGCCC ATTCTCCGCC 3061 CCATGGCTGA CTAATTTTTT TTATTTATGC AGAGGCCGAG GCCGCCTCGG CCTCTGAGCT 3121 ATTCCAGAAG TAGTGAGGAG GCTTTTTTGG AGGCCTAGGC TTTTGCAAAG ATCGATCAAG 3181 AGACAGGATG AGGATCGTTT CGCATGATTG AACAAGATGG ATTGCACGCA GGTTCTCCGG 3241 CCGCTTGGGT GGAGAGGCTA TTCGGCTATG ACTGGGCACA ACAGACAATC GGCTGCTCTG 3301 ATGCCGCCGT GTTCCGGCTG TCAGCGCAGG GGCGCCCGGT TCTTTTTGTC AAGACCGACC 3361 TGTCCGGTGC CCTGAATGAA CTGCAAGACG AGGCAGCGCG GCTATCGTGG CTGGCCACGA 3421 CGGGCGTTCC TTGCGCAGCT GTGCTCGACG TTGTCACTGA AGCGGGAAGG GACTGGCTGC 3481 TATTGGGCGA AGTGCCGGGG CAGGATCTCC TGTCATCTCA CCTTGCTCCT GCCGAGAAAG 3541 TATCCATCAT GGCTGATGCA ATGCGGCGGC TGCATACGCT TGATCCGGCT ACCTGCCCAT 3601 TCGACCACCA AGCGAAACAT CGCATCGAGC GAGCACGTAC TCGGATGGAA GCCGGTCTTG 3661 TCGATCAGGA TGATCTGGAC GAAGAGCATC AGGGGCTCGC GCCAGCCGAA CTGTTCGCCA 3721 GGCTCAAGGC GAGCATGCCC GACGGCGAGG ATCTCGTCGT GACCCATGGC GATGCCTGCT 3781 TGCCGAATAT CATGGTGGAA AATGGCCGCT TTTCTGGATT CATCGACTGT GGCCGGCTGG 3841 GTGTGGCGGA CCGCTATCAG GACATAGCGT TGGCTACCCG TGATATTGCT GAAGAGCTTG 3901 GCGGCGAATG GGCTGACCGC TTCCTCGTGC TTTACGGTAT CGCCGCTCCC GATTCGCAGC 3961 GCATCGCCTT CTATCGCCTT CTTGACGAGT TCTTCTGAGC GGGACTCTGG GGTTCGAAAT 4021 GACCGACCAA GCGACGCCCA ACCTGCCATC ACGAGATTTC GATTCCACCG CCGCCTTCTA 4081 TGAAAGGTTG GGCTTCGGAA TCGTTTTCCG GGACGCCGGC TGGATGATCC TCCAGCGCGG 4141 GGATCTCATG CTGGAGTTCT TCGCCCACCC TAGGGGGAGG CTAACTGAAA CACGGAAGGA 4201 GACAATACCG GAAGGAACCC GCGCTATGAC GGCAATAAAA AGACAGAATA AAACGCACGG 4261 TGTTGGGTCG TTTGTTCATA AACGCGGGGT TCGGTCCCAG GGCTGGCACT CTGTCGATAC 4321 CCCACCGAGA CCCCATTGGG GCCAATACGC CCGCGTTTCT TCCTTTTCCC CACCCCACCC 4381 CCCAAGTTCG GGTGAAGGCC CAGGGCTCGC AGCCAACGTC GGGGCGGCAG GCCCTGCCAT 4441 AGCCTCAGGT TACTCATATA TACTTTAGAT TGATTTAAAA CTTCATTTTT AATTTAAAAG 4501 GATCTAGGTG AAGATCCTTT TTGATAATCT CATGACCAAA ATCCCTTAAC GTGAGTTTTC 4561 GTTCCACTGA GCGTCAGACC CCGTAGAAAA GATCAAAGGA TCTTCTTGAG ATCCTTTTTT 4621 TCTGCGCGTA ATCTGCTGCT TGCAAACAAA AAAACCACCG CTACCAGCGG TGGTTTGTTT 4681 GCCGGATCAA GAGCTACCAA CTCTTTTTCC GAAGGTAACT GGCTTCAGCA GAGCGCAGAT 4741 ACCAAATACT GTCCTTCTAG TGTAGCCGTA GTTAGGCCAC CACTTCAAGA ACTCTGTAGC 4801 ACCGCCTACA TACCTCGCTC TGCTAATCCT GTTACCAGTG GCTGCTGCCA GTGGCGATAA 4861 GTCGTGTCTT ACCGGGTTGG ACTCAAGACG ATAGTTACCG GATAAGGCGC AGCGGTCGGG 4921 CTGAACGGGG GGTTCGTGCA CACAGCCCAG CTTGGAGCGA ACGACCTACA CCGAACTGAG 4981 ATACCTACAG CGTGAGCTAT GAGAAAGCGC CACGCTTCCC GAAGGGAGAA AGGCGGACAG 5041 GTATCCGGTA AGCGGCAGGG TCGGAACAGG AGAGCGCACG AGGGAGCTTC CAGGGGGAAA 5101 CGCCTGGTAT CTTTATAGTC CTGTCGGGTT TCGCCACCTC TGACTTGAGC GTCGATTTTT 5161 GTGATGCTCG TCAGGGGGGC GGAGCCTATG GAAAAACGCC AGCAACGCGG CCTTTTTACG 5221 GTTCCTGGCC TTTTGCTGGC CTTTTGCTCA CATGTTCTTT CCTGCGTTAT CCCCTGATTC 5281 TGTGGATAAC CGTATTACCG CCATGCAT //