| 出品公司: | Clontech |
|---|---|
| 载体名称: | pRetroQ-mCherry-C1 |
| 质粒类型: | 逆病毒载体;荧光报告载体 |
| 高拷贝/低拷贝: | 低拷贝 |
| 克隆方法: | 限制性内切酶,多克隆位点 |
| 启动子: | CMV IE |
| 载体大小: | 7605 bp |
| 5' 测序引物及序列: | -- |
| 3' 测序引物及序列: | -- |
| 载体标签: | mCherry(N-端) |
| 载体抗性: | 氨苄青霉素 |
| 筛选标记: | 嘌呤霉素(Puromycin) |
| 克隆菌株: | DH5α |
| 宿主细胞(系): | 常规细胞系(293、CV-1、CHO等),原代细胞 |
| 备注: |
pRetroQ-mCherry-C1载体是自我失活型逆病毒载体,表达N端mCherry融合蛋白; pRetroQ-mCherry-C1载体能够高效递送目的基因到靶细胞中,尤其适用于原代细胞和其 它难转染的细胞系。 |
| 产品目录号: | 632567 |
| 稳定性: | 稳表达 |
| 组成型/诱导型: | 组成型 |
| 病毒/非病毒: | 逆转录病毒 |
买家导航

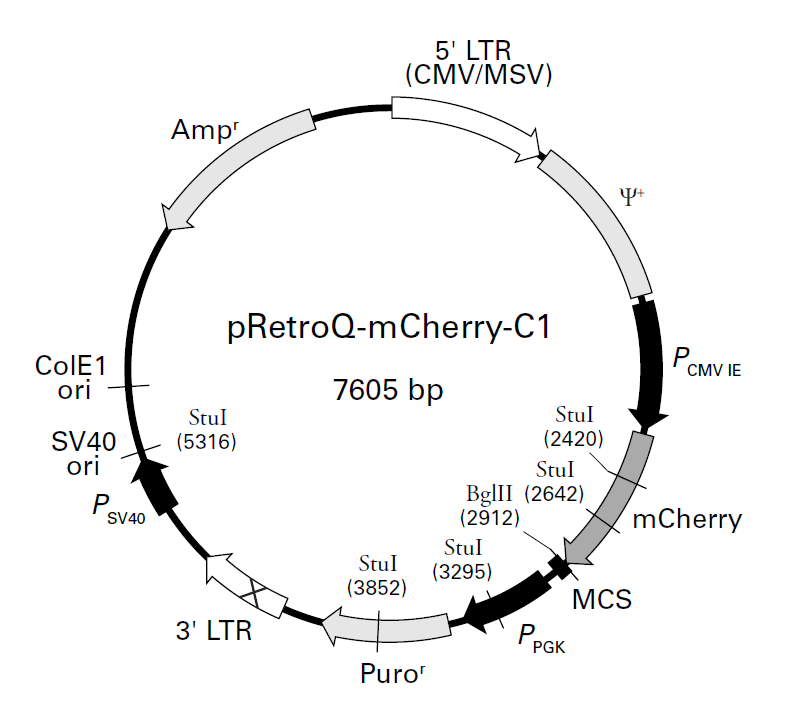
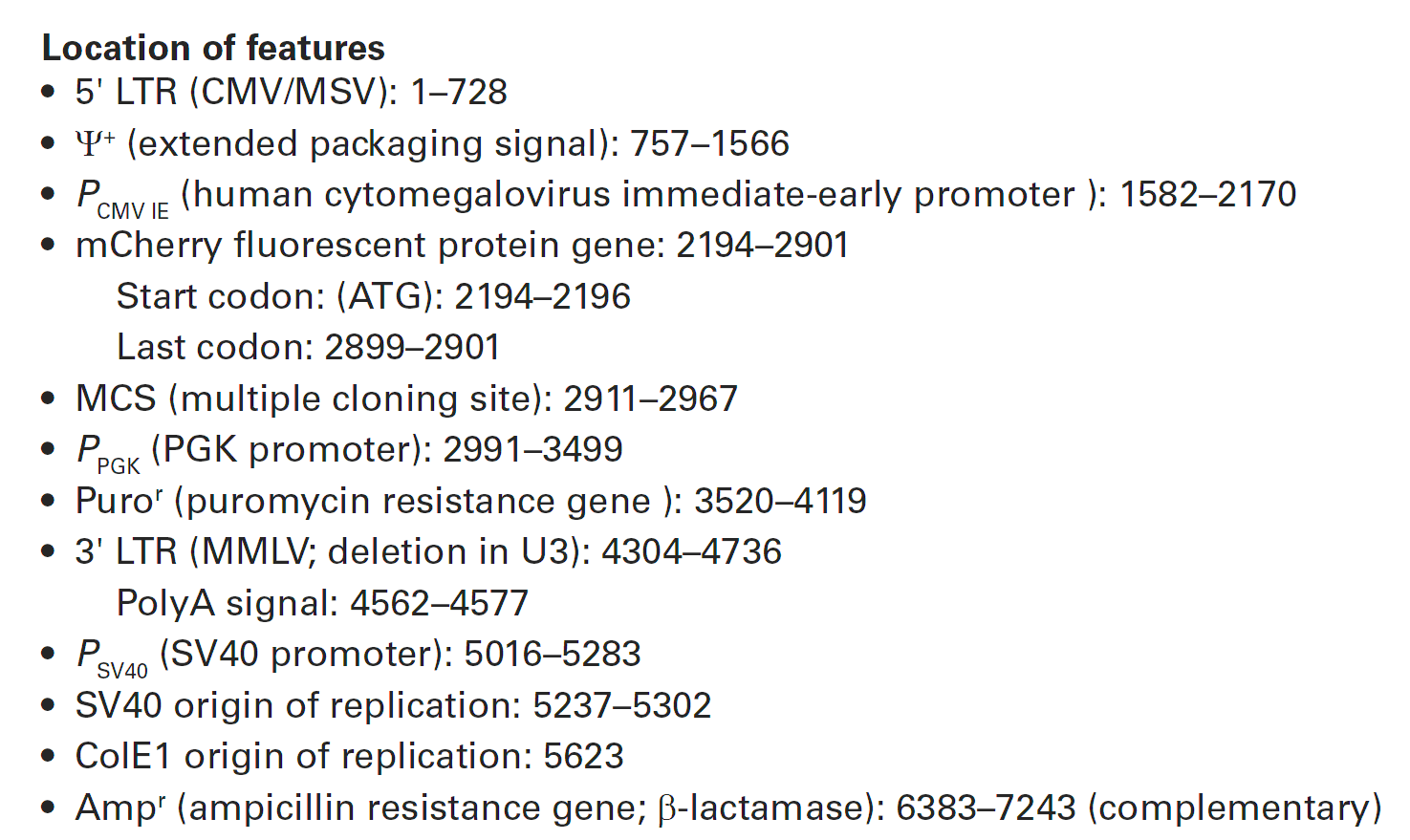
pRetroQ-mCherry-C1载体质粒基本信息
pRetroQ-mCherry-C1质粒图谱载体图谱和pRetroQ-mCherry-C1载体序列质粒序列多克隆位点信息
pRetroQ-mCherry-C1质粒载体简介
pRetroQ-mCherry-C1载体描述
pRetroQ-mCherry-C1 is a high-titer, self-inactivating retroviral vector that facilitates efficient delivery and expression of mCherry, or C-terminal mCherry fusions, to target cells. mCherry is a mutant fluorescent protein derived from the tetrameric Discosoma sp. red fluorescent protein, DsRed (1). The excitation and emission maxima of the native mCherry protein are 587 nm and 610 nm, respectively.
The multiple cloning site (MCS) in pRetroQ-mCherry-C1 is located just downstream of the mCherry coding sequence. Genes cloned into the MCS are expressed as C-terminal mCherry fusion proteins when they are in the same reading frame as mCherry and there are no intervening stop codons.
The RetroQ retroviral vector backbone incorporates several unique features. This vector contains a puromycin resistance cassette (Puror) driven by the PGK promoter (PPGK) for selection of positively-infected cells (2).The hybrid 5’ long terminal repeats (LTR) consists of the CMV type I enhancer and the murine sarcoma virus (MSV) promoter. This vector demonstrates high levels of transcription in HEK 293-based packaging cell lines due, in part, to the presence of adenoviral E1A (3–6) in these cells. The self-inactivating feature of the vector is provided by a deletion in the 3’ LTR enhancer region (U3). During reverse transcription of the retroviral RNA, the inactivated 3’ LTR is copied and replaces the 5’ LTR CMV enhancer sequences. This can reduce the phenomenon known as promoter interference (7) and allow more efficient expression.
Additionally, the viral genomic transcript contains the necessary viral RNA processing elements, including the LTRs, packaging signal (ψ+), and tRNA primer-binding site. pRetroQ-mCherry-C1 contains a bacterial origin of replication, an E. coli Ampr gene for propagation and selection in bacteria, and an SV40 origin for replication in mammalian cells expressing the SV40 large T antigen.
pRetroQ-mCherry-C1 is designed to efficiently deliver and express C-terminal mCherry fusions into primary cells or cells that are difficult to transfect. C-terminal mCherry fusion proteins retain the fluorescent properties of the native protein, allowing the in vivo localization of the fusion protein. The gene of interest should be cloned into pRetroQ-mCherry-C1 so that it is in frame with the mCherry coding sequence. The inserted sequence does not require an initiation codon (ATG) or a stop codon (TAA, TAG, TGA); however, if you don't want to use the stop codons downstream of the MCS (see map), you can add a stop codon to the end of your gene of interest. The recombinant mCherry vector can be transduced or transfected into mammalian cells. If required, stable transformants can be selected using puromycin. pRetroQ-mCherry-C1 can also be used simply to express mCherry in a cell line of interest (e.g., as an infection marker).
Before pRetroQ-mCherry-C1 can be transduced into mammalian cells, it must be transfected into a packaging cell line (such as the RetroPack PT67 Cell line, AmphoPack-293, EcoPack2-293, Pantropic Expression System , or Retro-X Universal Packaging System. The packaging cell line supplies the viral structural genes (gag, pol, and env) necessary for particle formation and replication that pRetroQ-mCherry-C1 lacks, allowing RNA from the vector to be packaged into non-infectious, replication-incompetent retroviral particles. Once a high-titer supernatant is produced, these retroviral particles can infect target cells and transmit the gene of interest, but they cannot replicate within the target cells due to the absence of viral structural genes. The separate introduction and integration of the structural genes into the packaging cell line minimizes the chances of producing replication-competent virus caused by recombination events during cell proliferation.
Propagation in E. coli
Suitable host strains: DH5α, Fusion Blue, and other general purpose strains.
Selectable marker: plasmid confers resistance to ampicillin (100 μg/ml) in E. coli hosts.
E. coli replication origin: ColE1
Copy number: low
Excitation and emission maxima of mCherry
Excitation maximum = 587 nm
Emission maximum = 610 nm
pRetroQ-mCherry-C1质粒序列载体序列
LOCUS pRetroQ-mCherry-C1 7605 bp DNA SYN
DEFINITION pRetroQ-mCherry-C1
ACCESSION
KEYWORDS
SOURCE
ORGANISM other sequences; artificial sequences; vectors.
FEATURES Location/Qualifiers
source 1..7605
/organism="pRetroQ-mCherry-C1"
/mol_type="other DNA"
promoter 2..506
/label="CMV_immearly_promoter"
misc_feature 6..688
/label="5_LTR"
misc_feature 9..296
/label="CAG_enhancer"
misc_feature 463..483
/label="CMV_fwd_primer"
misc_feature 758..1567
/label="psi_plus_pack"
misc_feature 1154..1561
/label="gag"
misc_feature 1497..1519
/label="MSCV_primer"
misc_feature 1497..1519
/label="pLXSN_5_primer"
misc_feature 1535..1551
/label="pBABE_5_primer"
promoter 1591..2143
/label="CMV_immearly_promoter"
misc_feature 1646..1933
/label="CAG_enhancer"
misc_feature 2100..2120
/label="CMV_fwd_primer"
promoter 2101..2170
/label="CMV_promoter"
CDS 2194..2982
/label="ORF frame 1"
/translation="MVSKGEEDNMAIIKEFMRFKVHMEGSVNGHEFEIEGEGEGRPYE
GTQTAKLKVTKGGPLPFAWDILSPQFMYGSKAYVKHPADIPDYLKLSFPEGFKWERVM
NFEDGGVVTVTQDSSLQDGEFIYKVKLRGTNFPSDGPVMQKKTMGWEASSERMYPEDG
ALKGEIKQRLKLKDGGHYDAEVKTTYKAKKPVQLPGAYNVNIKLDITSHNEDYTIVEQ
YERAEGRHSTGGMDELYKSGLRSRAQASNSAVDGTAGPGSTGSR*"
gene 2197..2901
/label="mCherry"
/gene="mCherry"
/translation="MVSKGEEDNMAIIKEFMRFKVHMEGSVNGHEFEIEGEGEGRPYE
GTQTAKLKVTKGGPLPFAWDILSPQFMYGSKAYVKHPADIPDYLKLSFPEGFKWERVM
NFEDGGVVTVTQDSSLQDGEFIYKVKLRGTNFPSDGPVMQKKTMGWEASSERMYPEDG
ALKGEIKQRLKLKDGGHYDAEVKTTYKAKKPVQLPGAYNVNIKLDITSHNEDYTIVEQ
YERAEGRHSTGGMDELYKSGLRSRAQASNSAVDGTAGPGSTGSR*"
misc_feature 2374..2427
/label="Q66M (mCherry)"
/translation="MVSKGEEDNMAIIKEFMRFKVHMEGSVNGHEFEIEGEGEGRPYE
GTQTAKLKVTKGGPLPFAWDILSPQFMYGSKAYVKHPADIPDYLKLSFPEGFKWERVM
NFEDGGVVTVTQDSSLQDGEFIYKVKLRGTNFPSDGPVMQKKTMGWEASSERMYPEDG
ALKGEIKQRLKLKDGGHYDAEVKTTYKAKKPVQLPGAYNVNIKLDITSHNEDYTIVEQ
YERAEGRHSTGGMDELYKSGLRSRAQASNSAVDGTAGPGSTGSR*"
misc_feature complement(3035..3058)
/label="MSCV_rev_primer"
/translation="MVSKGEEDNMAIIKEFMRFKVHMEGSVNGHEFEIEGEGEGRPYE
GTQTAKLKVTKGGPLPFAWDILSPQFMYGSKAYVKHPADIPDYLKLSFPEGFKWERVM
NFEDGGVVTVTQDSSLQDGEFIYKVKLRGTNFPSDGPVMQKKTMGWEASSERMYPEDG
ALKGEIKQRLKLKDGGHYDAEVKTTYKAKKPVQLPGAYNVNIKLDITSHNEDYTIVEQ
YERAEGRHSTGGMDELYKSGLRSRAQASNSAVDGTAGPGSTGSR*"
CDS 3280..4119
/label="ORF frame 1"
/translation="MEAGRPLGQRPIAALLLRFLGSEAGKGWVRGRAQGRAQGRGGRP
KVLRRPGILHASKAHVCRAVLLFLISGPFDLQPKLTMTEYKPTVRLATRDDVPRAVRT
LAAAFADYPATRHTVDPDRHIERVTELQELFLTRVGLDIGKVWVADDGAAVAVWTTPE
SVEAGAVFAEIGPRMAELSGSRLAAQQQMEGLLAPHRPKEPAWFLATVGVSPDHQGKG
LGSAVVLPGVEAAERAGVPAFLETSAPRNLPFYERLGFTVTADVEVPEGPRTWCMTRK
PGA*"
misc_feature 3432..3451
/label="mPGK_F_primer"
/translation="MEAGRPLGQRPIAALLLRFLGSEAGKGWVRGRAQGRAQGRGGRP
KVLRRPGILHASKAHVCRAVLLFLISGPFDLQPKLTMTEYKPTVRLATRDDVPRAVRT
LAAAFADYPATRHTVDPDRHIERVTELQELFLTRVGLDIGKVWVADDGAAVAVWTTPE
SVEAGAVFAEIGPRMAELSGSRLAAQQQMEGLLAPHRPKEPAWFLATVGVSPDHQGKG
LGSAVVLPGVEAAERAGVPAFLETSAPRNLPFYERLGFTVTADVEVPEGPRTWCMTRK
PGA*"
CDS complement(3445..4170)
/label="ORF frame 1"
/translation="MGSCAPFGRALRVVGRASGTGLAGHAPGARSFGHLDVGGDGEAE
PLVEGEVAGRGGLQEGGHPGALGRLHSGEHDGAAQTLALVVGRDADGGQEPRGLLGPV
RRQEAFHLLLRGQPGTAQLGHARADLGEHRPRFDALRRGPDRHRGAVVRDPHLADVEP
DAREEEFLQLGDPLDVAVRIDGVARGGVVGERGGEGAYGPGDVVAGGEAHRGLVLGHG
KLGLQVERPGDEEEENSAADVRF*"
gene 3520..4119
/label="puro (variant)"
/gene="puro (variant)"
/translation="MGSCAPFGRALRVVGRASGTGLAGHAPGARSFGHLDVGGDGEAE
PLVEGEVAGRGGLQEGGHPGALGRLHSGEHDGAAQTLALVVGRDADGGQEPRGLLGPV
RRQEAFHLLLRGQPGTAQLGHARADLGEHRPRFDALRRGPDRHRGAVVRDPHLADVEP
DAREEEFLQLGDPLDVAVRIDGVARGGVVGERGGEGAYGPGDVVAGGEAHRGLVLGHG
KLGLQVERPGDEEEENSAADVRF*"
misc_feature complement(4839..4861)
/label="pGEX_3_primer"
/translation="MGSCAPFGRALRVVGRASGTGLAGHAPGARSFGHLDVGGDGEAE
PLVEGEVAGRGGLQEGGHPGALGRLHSGEHDGAAQTLALVVGRDADGGQEPRGLLGPV
RRQEAFHLLLRGQPGTAQLGHARADLGEHRPRFDALRRGPDRHRGAVVRDPHLADVEP
DAREEEFLQLGDPLDVAVRIDGVARGGVVGERGGEGAYGPGDVVAGGEAHRGLVLGHG
KLGLQVERPGDEEEENSAADVRF*"
misc_feature complement(4997..5017)
/label="pBABE_3_primer"
/translation="MGSCAPFGRALRVVGRASGTGLAGHAPGARSFGHLDVGGDGEAE
PLVEGEVAGRGGLQEGGHPGALGRLHSGEHDGAAQTLALVVGRDADGGQEPRGLLGPV
RRQEAFHLLLRGQPGTAQLGHARADLGEHRPRFDALRRGPDRHRGAVVRDPHLADVEP
DAREEEFLQLGDPLDVAVRIDGVARGGVVGERGGEGAYGPGDVVAGGEAHRGLVLGHG
KLGLQVERPGDEEEENSAADVRF*"
misc_feature complement(5003..5218)
/label="SV40_enhancer"
/translation="MGSCAPFGRALRVVGRASGTGLAGHAPGARSFGHLDVGGDGEAE
PLVEGEVAGRGGLQEGGHPGALGRLHSGEHDGAAQTLALVVGRDADGGQEPRGLLGPV
RRQEAFHLLLRGQPGTAQLGHARADLGEHRPRFDALRRGPDRHRGAVVRDPHLADVEP
DAREEEFLQLGDPLDVAVRIDGVARGGVVGERGGEGAYGPGDVVAGGEAHRGLVLGHG
KLGLQVERPGDEEEENSAADVRF*"
promoter 5015..5283
/label="SV40_promoter"
/translation="MGSCAPFGRALRVVGRASGTGLAGHAPGARSFGHLDVGGDGEAE
PLVEGEVAGRGGLQEGGHPGALGRLHSGEHDGAAQTLALVVGRDADGGQEPRGLLGPV
RRQEAFHLLLRGQPGTAQLGHARADLGEHRPRFDALRRGPDRHRGAVVRDPHLADVEP
DAREEEFLQLGDPLDVAVRIDGVARGGVVGERGGEGAYGPGDVVAGGEAHRGLVLGHG
KLGLQVERPGDEEEENSAADVRF*"
rep_origin 5182..5259
/label="SV40_origin"
/translation="MGSCAPFGRALRVVGRASGTGLAGHAPGARSFGHLDVGGDGEAE
PLVEGEVAGRGGLQEGGHPGALGRLHSGEHDGAAQTLALVVGRDADGGQEPRGLLGPV
RRQEAFHLLLRGQPGTAQLGHARADLGEHRPRFDALRRGPDRHRGAVVRDPHLADVEP
DAREEEFLQLGDPLDVAVRIDGVARGGVVGERGGEGAYGPGDVVAGGEAHRGLVLGHG
KLGLQVERPGDEEEENSAADVRF*"
misc_feature 5244..5263
/label="SV40pro_F_primer"
/translation="MGSCAPFGRALRVVGRASGTGLAGHAPGARSFGHLDVGGDGEAE
PLVEGEVAGRGGLQEGGHPGALGRLHSGEHDGAAQTLALVVGRDADGGQEPRGLLGPV
RRQEAFHLLLRGQPGTAQLGHARADLGEHRPRFDALRRGPDRHRGAVVRDPHLADVEP
DAREEEFLQLGDPLDVAVRIDGVARGGVVGERGGEGAYGPGDVVAGGEAHRGLVLGHG
KLGLQVERPGDEEEENSAADVRF*"
gene complement(6383..7243)
/label="Ampicillin"
/gene="Ampicillin"
/translation="MGSCAPFGRALRVVGRASGTGLAGHAPGARSFGHLDVGGDGEAE
PLVEGEVAGRGGLQEGGHPGALGRLHSGEHDGAAQTLALVVGRDADGGQEPRGLLGPV
RRQEAFHLLLRGQPGTAQLGHARADLGEHRPRFDALRRGPDRHRGAVVRDPHLADVEP
DAREEEFLQLGDPLDVAVRIDGVARGGVVGERGGEGAYGPGDVVAGGEAHRGLVLGHG
KLGLQVERPGDEEEENSAADVRF*"
CDS complement(6383..7243)
/label="ORF frame 3"
/translation="MSIQHFRVALIPFFAAFCLPVFAHPETLVKVKDAEDQLGARVGY
IELDLNSGKILESFRPEERFPMMSTFKVLLCGAVLSRVDAGQEQLGRRIHYSQNDLVE
YSPVTEKHLTDGMTVRELCSAAITMSDNTAANLLLTTIGGPKELTAFLHNMGDHVTRL
DRWEPELNEAIPNDERDTTMPAAMATTLRKLLTGELLTLASRQQLIDWMEADKVAGPL
LRSALPAGWFIADKSGAGERGSRGIIAALGPDGKPSRIVVIYTTGSQATMDERNRQIA
EIGASLIKHW*"
promoter complement(7285..7313)
/label="AmpR_promoter"
/translation="MSIQHFRVALIPFFAAFCLPVFAHPETLVKVKDAEDQLGARVGY
IELDLNSGKILESFRPEERFPMMSTFKVLLCGAVLSRVDAGQEQLGRRIHYSQNDLVE
YSPVTEKHLTDGMTVRELCSAAITMSDNTAANLLLTTIGGPKELTAFLHNMGDHVTRL
DRWEPELNEAIPNDERDTTMPAAMATTLRKLLTGELLTLASRQQLIDWMEADKVAGPL
LRSALPAGWFIADKSGAGERGSRGIIAALGPDGKPSRIVVIYTTGSQATMDERNRQIA
EIGASLIKHW*"
promoter 7535..506
/label="CMV_immearly_promoter"
/translation="MSIQHFRVALIPFFAAFCLPVFAHPETLVKVKDAEDQLGARVGY
IELDLNSGKILESFRPEERFPMMSTFKVLLCGAVLSRVDAGQEQLGRRIHYSQNDLVE
YSPVTEKHLTDGMTVRELCSAAITMSDNTAANLLLTTIGGPKELTAFLHNMGDHVTRL
DRWEPELNEAIPNDERDTTMPAAMATTLRKLLTGELLTLASRQQLIDWMEADKVAGPL
LRSALPAGWFIADKSGAGERGSRGIIAALGPDGKPSRIVVIYTTGSQATMDERNRQIA
EIGASLIKHW*"
misc_feature 7599..688
/label="5_LTR"
/translation="MSIQHFRVALIPFFAAFCLPVFAHPETLVKVKDAEDQLGARVGY
IELDLNSGKILESFRPEERFPMMSTFKVLLCGAVLSRVDAGQEQLGRRIHYSQNDLVE
YSPVTEKHLTDGMTVRELCSAAITMSDNTAANLLLTTIGGPKELTAFLHNMGDHVTRL
DRWEPELNEAIPNDERDTTMPAAMATTLRKLLTGELLTLASRQQLIDWMEADKVAGPL
LRSALPAGWFIADKSGAGERGSRGIIAALGPDGKPSRIVVIYTTGSQATMDERNRQIA
EIGASLIKHW*"
ORIGIN
1 TCCGCGTTAC ATAACTTACG GTAAATGGCC CGCCTGGCTG ACCGCCCAAC GACCCCCGCC
61 CATTGACGTC AATAATGACG TATGTTCCCA TAGTAACGCC AATAGGGACT TTCCATTGAC
121 GTCAATGGGT GGAGTATTTA CGGTAAACTG CCCACTTGGC AGTACATCAA GTGTATCATA
181 TGCCAAGTAC GCCCCCTATT GACGTCAATG ACGGTAAATG GCCCGCCTGG CATTATGCCC
241 AGTACATGAC CTTATGGGAC TTTCCTACTT GGCAGTACAT CTACGTATTA GTCATCGCTA
301 TTACCATGGT GATGCGGTTT TGGCAGTACA TCAATGGGCG TGGATAGCGG TTTGACTCAC
361 GGGGATTTCC AAGTCTCCAC CCCATTGACG TCAATGGGAG TTTGTTTTGG CACCAAAATC
421 AACGGGACTT TCCAAAATGT CGTAACAACT CCGCCCCATT GACGCAAATG GGCGGTAGGC
481 GTGTACGGTG GGAGGTCTAT ATAAGCAGAG CTCAATAAAA GAGCCCACAA CCCCTCACTC
541 GGCGCGCCAG TCTTCCGATA GACTGCGTCG CCCGGGTACC CGTATTCCCA ATAAAGCCTC
601 TTGCTGTTTG CATCCGAATC GTGGTCTCGC TGTTCCTTGG GAGGGTCTCC TCTGAGTGAT
661 TGACTACCCA CGACGGGGGT CTTTCATTTG GGGGCTCGTC CGGGATTTGG AGACCCCTGC
721 CCAGGGACCA CCGACCCACC ACCGGGAGGT AAGCTGGCCA GCAACTTATC TGTGTCTGTC
781 CGATTGTCTA GTGTCTATGT TTGATGTTAT GCGCCTGCGT CTGTACTAGT TAGCTAACTA
841 GCTCTGTATC TGGCGGACCC GTGGTGGAAC TGACGAGTTC TGAACACCCG GCCGCAACCC
901 TGGGAGACGT CCCAGGGACT TTGGGGGCCG TTTTTGTGGC CCGACCTGAG GAAGGGAGTC
961 GATGTGGAAT CCGACCCCGT CAGGATATGT GGTTCTGGTA GGAGACGAGA ACCTAAAACA
1021 GTTCCCGCCT CCGTCTGAAT TTTTGCTTTC GGTTTGGAAC CGAAGCCGCG CGTCTTGTCT
1081 GCTGCAGCGC TGCAGCATCG TTCTGTGTTG TCTCTGTCTG ACTGTGTTTC TGTATTTGTC
1141 TGAAAATTAG GGCCAGACTG TTACCACTCC CTTAAGTTTG ACCTTAGGTC ACTGGAAAGA
1201 TGTCGAGCGG ATCGCTCACA ACCAGTCGGT AGATGTCAAG AAGAGACGTT GGGTTACCTT
1261 CTGCTCTGCA GAATGGCCAA CCTTTAACGT CGGATGGCCG CGAGACGGCA CCTTTAACCG
1321 AGACCTCATC ACCCAGGTTA AGATCAAGGT CTTTTCACCT GGCCCGCATG GACACCCAGA
1381 CCAGGTCCCC TACATCGTGA CCTGGGAAGC CTTGGCTTTT GACCCCCCTC CCTGGGTCAA
1441 GCCCTTTGTA CACCCTAAGC CTCCGCCTCC TCTTCCTCCA TCCGCCCCGT CTCTCCCCCT
1501 TGAACCTCCT CGTTCGACCC CGCCTCGATC CTCCCTTTAT CCAGCCCTCA CTCCTTCTCT
1561 AGGCGCCGGA ATTGAAGATC ATAGTTATTA ATAGTAATCA ATTACGGGGT CATTAGTTCA
1621 TAGCCCATAT ATGGAGTTCC GCGTTACATA ACTTACGGTA AATGGCCCGC CTGGCTGACC
1681 GCCCAACGAC CCCCGCCCAT TGACGTCAAT AATGACGTAT GTTCCCATAG TAACGCCAAT
1741 AGGGACTTTC CATTGACGTC AATGGGTGGA GTATTTACGG TAAACTGCCC ACTTGGCAGT
1801 ACATCAAGTG TATCATATGC CAAGTACGCC CCCTATTGAC GTCAATGACG GTAAATGGCC
1861 CGCCTGGCAT TATGCCCAGT ACATGACCTT ATGGGACTTT CCTACTTGGC AGTACATCTA
1921 CGTATTAGTC ATCGCTATTA CCATGGTGAT GCGGTTTTGG CAGTACATCA ATGGGCGTGG
1981 ATAGCGGTTT GACTCACGGG GATTTCCAAG TCTCCACCCC ATTGACGTCA ATGGGAGTTT
2041 GTTTTGGCAC CAAAATCAAC GGGACTTTCC AAAATGTCGT AACAACTCCG CCCCATTGAC
2101 GCAAATGGGC GGTAGGCGTG TACGGTGGGA GGTCTATATA AGCAGAGCTG GTTTAGTGAA
2161 CCGTCAGATC CGCTAGCGCT ACCGGTCGCC ACCATGGTGA GCAAGGGCGA GGAGGATAAC
2221 ATGGCCATCA TCAAGGAGTT CATGCGCTTC AAGGTGCACA TGGAGGGCTC CGTGAACGGC
2281 CACGAGTTCG AGATCGAGGG CGAGGGCGAG GGCCGCCCCT ACGAGGGCAC CCAGACCGCC
2341 AAGCTGAAGG TGACCAAGGG TGGCCCCCTG CCCTTCGCCT GGGACATCCT GTCCCCTCAG
2401 TTCATGTACG GCTCCAAGGC CTACGTGAAG CACCCCGCCG ACATCCCCGA CTACTTGAAG
2461 CTGTCCTTCC CCGAGGGCTT CAAGTGGGAG CGCGTGATGA ACTTCGAGGA CGGCGGCGTG
2521 GTGACCGTGA CCCAGGACTC CTCCCTGCAG GACGGCGAGT TCATCTACAA GGTGAAGCTG
2581 CGCGGCACCA ACTTCCCCTC CGACGGCCCC GTAATGCAGA AGAAGACCAT GGGCTGGGAG
2641 GCCTCCTCCG AGCGGATGTA CCCCGAGGAC GGCGCCCTGA AGGGCGAGAT CAAGCAGAGG
2701 CTGAAGCTGA AGGACGGCGG CCACTACGAC GCTGAGGTCA AGACCACCTA CAAGGCCAAG
2761 AAGCCCGTGC AGCTGCCCGG CGCCTACAAC GTCAACATCA AGTTGGACAT CACCTCCCAC
2821 AACGAGGACT ACACCATCGT GGAACAGTAC GAACGCGCCG AGGGCCGCCA CTCCACCGGC
2881 GGCATGGACG AGCTGTACAA GTCCGGACTC AGATCTCGAG CTCAAGCTTC GAATTCTGCA
2941 GTCGACGGTA CCGCGGGCCC GGGATCCACC GGATCTAGAT AACTGATCAT AATTCTACCG
3001 GGTAGGGGAG GCGCTTTTCC CAAGGCAGTC TGGAGCATGC GCTTTAGCAG CCCCGCTGGG
3061 CACTTGGCGC TACACAAGTG GCCTCTGGCC TCGCACACAT TCCACATCCA CCGGTAGGCG
3121 CCAACCGGCT CCGTTCTTTG GTGGCCCCTT CGCGCCACCT TCTACTCCTC CCCTAGTCAG
3181 GAAGTTCCCC CCCGCCCCGC AGCTCGCGTC GTGCAGGACG TGACAAATGG AAGTAGCACG
3241 TCTCACTAGT CTCGTGCAGA TGGACAGCAC CGCTGAGCAA TGGAAGCGGG TAGGCCTTTG
3301 GGGCAGCGGC CAATAGCAGC TTTGCTCCTT CGCTTTCTGG GCTCAGAGGC TGGGAAGGGG
3361 TGGGTCCGGG GGCGGGCTCA GGGGCGGGCT CAGGGGCGGG GCGGGCGCCC GAAGGTCCTC
3421 CGGAGGCCCG GCATTCTGCA CGCTTCAAAA GCGCACGTCT GCCGCGCTGT TCTCCTCTTC
3481 CTCATCTCCG GGCCTTTCGA CCTGCAGCCC AAGCTTACCA TGACCGAGTA CAAGCCCACG
3541 GTGCGCCTCG CCACCCGCGA CGACGTCCCC AGGGCCGTAC GCACCCTCGC CGCCGCGTTC
3601 GCCGACTACC CCGCCACGCG CCACACCGTC GATCCGGACC GCCACATCGA GCGGGTCACC
3661 GAGCTGCAAG AACTCTTCCT CACGCGCGTC GGGCTCGACA TCGGCAAGGT GTGGGTCGCG
3721 GACGACGGCG CCGCGGTGGC GGTCTGGACC ACGCCGGAGA GCGTCGAAGC GGGGGCGGTG
3781 TTCGCCGAGA TCGGCCCGCG CATGGCCGAG TTGAGCGGTT CCCGGCTGGC CGCGCAGCAA
3841 CAGATGGAAG GCCTCCTGGC GCCGCACCGG CCCAAGGAGC CCGCGTGGTT CCTGGCCACC
3901 GTCGGCGTCT CGCCCGACCA CCAGGGCAAG GGTCTGGGCA GCGCCGTCGT GCTCCCCGGA
3961 GTGGAGGCGG CCGAGCGCGC CGGGGTGCCC GCCTTCCTGG AGACCTCCGC GCCCCGCAAC
4021 CTCCCCTTCT ACGAGCGGCT CGGCTTCACC GTCACCGCCG ACGTCGAGGT GCCCGAAGGA
4081 CCGCGCACCT GGTGCATGAC CCGCAAGCCC GGTGCCTGAC GCCCGCCCCA CGACCCGCAG
4141 CGCCCGACCG AAAGGAGCGC ACGACCCCAT GCATCGGCGT CTCGATCGAG ATATCAGTGG
4201 TCCAGGCTCT AGTTTTGACT CAACAATATC ACCAGCTGAA GCCTATAGAG TACGAGCCAT
4261 AGATAAAATA AAAGATTTTA TTTAGTCTCC AGAAAAAGGG GGGAATGAAA GACCCCACCT
4321 GTAGGTTTGG CAAGCTACTA GCTTAAGTAA CGCCATTTTG CAAGGCATGG AAAAATACAT
4381 AACTGAGAAT AGAGAAGTTC AGATCAAGGT CAGGAACAGA TGGAACAGGG TCGATCGACC
4441 CTAGAGAACC ATCAGATGTT TCCAGGGTGC CCCAAGGACC TGAAATGACC CTGTGCCTTA
4501 TTTGAACTAA CCAATCAGTT CGCTTCTCGC TTCTGTTCGC GCGCTTCTGC TCCCCGAGCT
4561 CAATAAAAGA GCCCACAACC CCTCACTCGG GGCGCCAGTC CTCCGATTGA CTGAGTCGCC
4621 CGGGTACCCG TGTATCCAAT AAACCCTCTT GCAGTTGCAT CCGACTTGTG GTCTCGCTGT
4681 TCCTTGGGAG GGTCTCCTCT GAGTGATTGA CTACCCGTCA GCGGGGGTCT TTCATTTGGG
4741 GGCTCGTCCG GGATCGGGAG ACCCCTGCCC AGGGACCACC GACCCACCAC CGGGAGGTAA
4801 GCTGGCTGCC TCGCGCGTTT CGGTGATGAC GGTGAAAACC TCTGACACAT GCAGCTCCCG
4861 GAGACGGTCA CAGCTTGTCT GTAAGCGGAT GCCGGGAGCA GACAAGCCCG TCAGGGCGCG
4921 TCAGCGGGTG TTGGCGGGTG TCGGGGCGCA GCCATGACCC AGTCACGTAG CGATAGCGGA
4981 GTGTAGATCC GGCTGTGGAA TGTGTGTCAG TTAGGGTGTG GAAAGTCCCC AGGCTCCCCA
5041 GCAGGCAGAA GTATGCAAAG CATGCATCTC AATTAGTCAG CAACCAGGTG TGGAAAGTCC
5101 CCAGGCTCCC CAGCAGGCAG AAGTATGCAA AGCATGCATC TCAATTAGTC AGCAACCATA
5161 GTCCCGCCCC TAACTCCGCC CATCCCGCCC CTAACTCCGC CCAGTTCCGC CCATTCTCCG
5221 CCCCATGGCT GACTAATTTT TTTTATTTAT GCAGAGGCCG AGGCCGCCTC GGCCTCTGAG
5281 CTATTCCAGA AGTAGTGAGG AGGCTTTTTT GGAGGCCTAG GCTTTTGCAA AAAGCTTACT
5341 GGCTTAACTA TGCGGCATCA GAGCAGATTG TACTGAGAGT GCACCATATG CGGTGTGAAA
5401 TACCGCACAG ATGCGTAAGG AGAAAATACC GCATCAGGCG CTCTTCCGCT TCCTCGCTCA
5461 CTGACTCGCT GCGCTCGGTC GTTCGGCTGC GGCGAGCGGT ATCAGCTCAC TCAAAGGCGG
5521 TAATACGGTT ATCCACAGAA TCAGGGGATA ACGCAGGAAA GAACATGTGA GCAAAAGGCC
5581 AGCAAAAGGC CAGGAACCGT AAAAAGGCCG CGTTGCTGGC GTTTTTCCAT AGGCTCCGCC
5641 CCCCTGACGA GCATCACAAA AATCGACGCT CAAGTCAGAG GTGGCGAAAC CCGACAGGAC
5701 TATAAAGATA CCAGGCGTTT CCCCCTGGAA GCTCCCTCGT GCGCTCTCCT GTTCCGACCC
5761 TGCCGCTTAC CGGATACCTG TCCGCCTTTC TCCCTTCGGG AAGCGTGGCG CTTTCTCATA
5821 GCTCACGCTG TAGGTATCTC AGTTCGGTGT AGGTCGTTCG CTCCAAGCTG GGCTGTGTGC
5881 ACGAACCCCC CGTTCAGCCC GACCGCTGCG CCTTATCCGG TAACTATCGT CTTGAGTCCA
5941 ACCCGGTAAG ACACGACTTA TCGCCACTGG CAGCAGCCAC TGGTAACAGG ATTAGCAGAG
6001 CGAGGTATGT AGGCGGTGCT ACAGAGTTCT TGAAGTGGTG GCCTAACTAC GGCTACACTA
6061 GAAGGACAGT ATTTGGTATC TGCGCTCTGC TGAAGCCAGT TACCTTCGGA AAAAGAGTTG
6121 GTAGCTCTTG ATCCGGCAAA CAAACCACCG CTGGTAGCGG TGGTTTTTTT GTTTGCAAGC
6181 AGCAGATTAC GCGCAGAAAA AAAGGATCTC AAGAAGATCC TTTGATCTTT TCTACGGGGT
6241 CTGACGCTCA GTGGAACGAA AACTCACGTT AAGGGATTTT GGTCATGAGA TTATCAAAAA
6301 GGATCTTCAC CTAGATCCTT TTAAATTAAA AATGAAGTTT TAAATCAATC TAAAGTATAT
6361 ATGAGTAAAC TTGGTCTGAC AGTTACCAAT GCTTAATCAG TGAGGCACCT ATCTCAGCGA
6421 TCTGTCTATT TCGTTCATCC ATAGTTGCCT GACTCCCCGT CGTGTAGATA ACTACGATAC
6481 GGGAGGGCTT ACCATCTGGC CCCAGTGCTG CAATGATACC GCGAGACCCA CGCTCACCGG
6541 CTCCAGATTT ATCAGCAATA AACCAGCCAG CCGGAAGGGC CGAGCGCAGA AGTGGTCCTG
6601 CAACTTTATC CGCCTCCATC CAGTCTATTA ATTGTTGCCG GGAAGCTAGA GTAAGTAGTT
6661 CGCCAGTTAA TAGTTTGCGC AACGTTGTTG CCATTGCTGC AGGCATCGTG GTGTCACGCT
6721 CGTCGTTTGG TATGGCTTCA TTCAGCTCCG GTTCCCAACG ATCAAGGCGA GTTACATGAT
6781 CCCCCATGTT GTGCAAAAAA GCGGTTAGCT CCTTCGGTCC TCCGATCGTT GTCAGAAGTA
6841 AGTTGGCCGC AGTGTTATCA CTCATGGTTA TGGCAGCACT GCATAATTCT CTTACTGTCA
6901 TGCCATCCGT AAGATGCTTT TCTGTGACTG GTGAGTACTC AACCAAGTCA TTCTGAGAAT
6961 AGTGTATGCG GCGACCGAGT TGCTCTTGCC CGGCGTCAAC ACGGGATAAT ACCGCGCCAC
7021 ATAGCAGAAC TTTAAAAGTG CTCATCATTG GAAAACGTTC TTCGGGGCGA AAACTCTCAA
7081 GGATCTTACC GCTGTTGAGA TCCAGTTCGA TGTAACCCAC TCGTGCACCC AACTGATCTT
7141 CAGCATCTTT TACTTTCACC AGCGTTTCTG GGTGAGCAAA AACAGGAAGG CAAAATGCCG
7201 CAAAAAAGGG AATAAGGGCG ACACGGAAAT GTTGAATACT CATACTCTTC CTTTTTCAAT
7261 ATTATTGAAG CATTTATCAG GGTTATTGTC TCATGAGCGG ATACATATTT GAATGTATTT
7321 AGAAAAATAA ACAAATAGGG GTTCCGCGCA CATTTCCCCG AAAAGTGCCA CCTGACGTCT
7381 AAGAAACCAT TATTATCATG ACATTAACCT ATAAAAATAG GCGTATCACG AGGCCCTTTC
7441 GTCTTCAAGA ATTAGCTTGG CCATTGCATA CGTTGTATCC ATATCATAAT ATGTACATTT
7501 ATATTGGCTC ATGTCCAACA TTACCGCCAT GTTGACATTG ATTATTGACT AGTTATTAAT
7561 AGTAATCAAT TACGGGGTCA TTAGTTCATA GCCCATATAT GGAGT
//