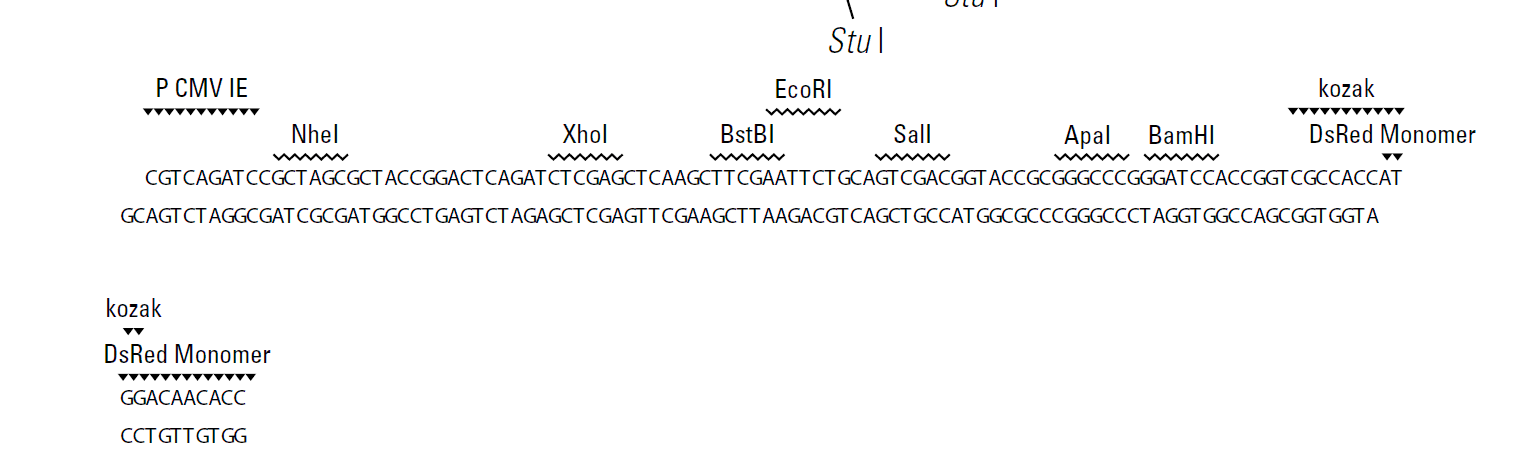| 出品公司: | Clontech |
|---|---|
| 载体名称: | pRetroQ-DsRed-Monomer-N1 |
| 质粒类型: | 逆病毒载体;荧光报告载体 |
| 高拷贝/低拷贝: | 低拷贝 |
| 克隆方法: | 限制性内切酶,多克隆位点 |
| 启动子: | CMV IE |
| 载体大小: | 7571 bp |
| 5' 测序引物及序列: | -- |
| 3' 测序引物及序列: | -- |
| 载体标签: | DsRed-Monomer(C-端) |
| 载体抗性: | 氨苄青霉素 |
| 筛选标记: | 嘌呤霉素(Puromycin) |
| 克隆菌株: | DH5α |
| 宿主细胞(系): | 常规细胞系(293、CV-1、CHO等),原代细胞 |
| 备注: |
pRetroQ-DsRed-Monomer-N1载体是自我失活型逆病毒载体,表达C端DsRed-Monomer融合蛋白; pRetroQ-DsRed-Monomer-N1载体能够高效递送目的基因到靶细胞中,尤其适用于原代细胞和其它难转染的细胞系; DsRed-Monomer是DsRed的单体突变体,与DsRed相比,编码序列经过了人工优化,适用于在哺乳动物细胞中的高水平表达。 |
| 产品目录号: | 632507 |
| 稳定性: | 稳表达 |
| 组成型/诱导型: | 组成型 |
| 病毒/非病毒: | 逆转录病毒 |
买家导航

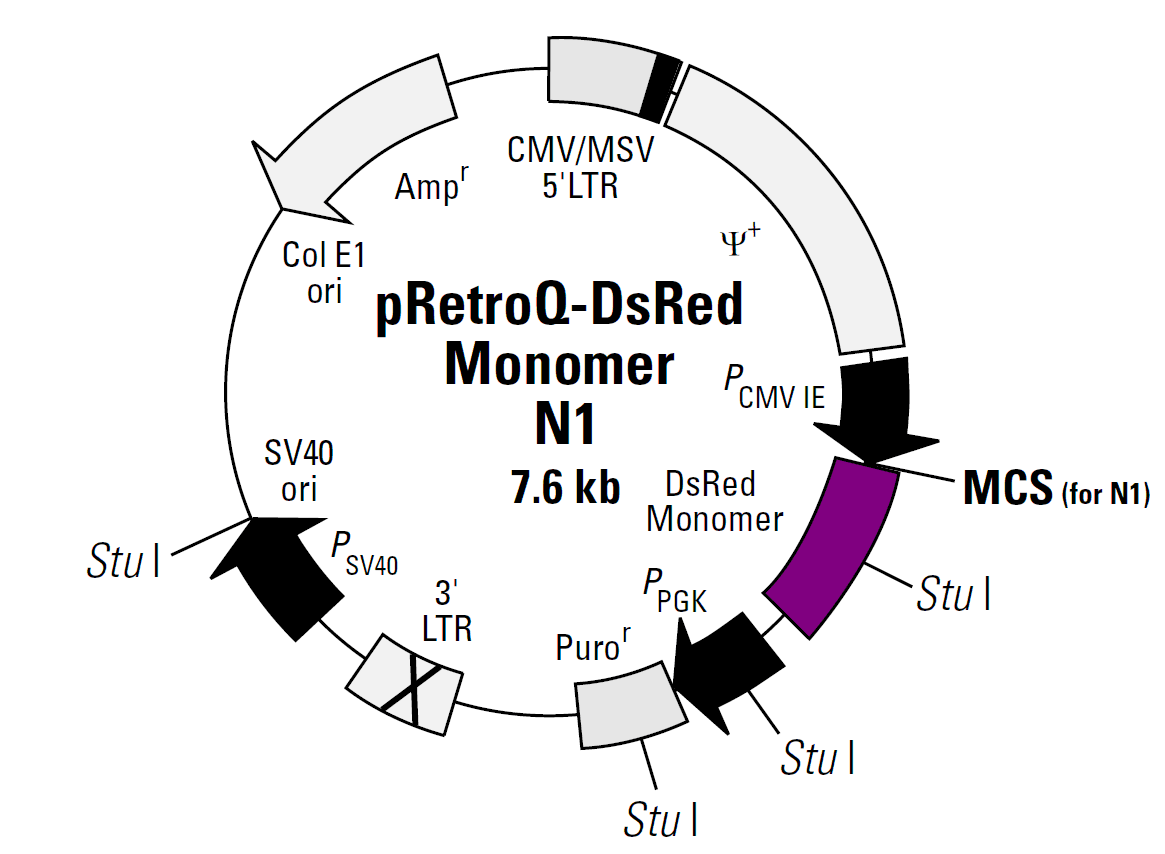
pRetroQ-DsRed-Monomer-N1载体质粒基本信息
pRetroQ-DsRed-Monomer-N1质粒图谱载体图谱和pRetroQ-DsRed-Monomer-N1载体序列质粒序列多克隆位点信息
pRetroQ-DsRed-Monomer-N1质粒载体简介
pRetroQ-DsRed Monomer-N1载体描述
pRetroQ-DsRed Monomer-N1 is a high-titer, self-inactivating retroviral vector that facilitates efficient delivery and expression of DsRed-monomer (DsRed.M1) as well as N terminal fusions of DsRed monomer to target cells. DsRed.M1 is a monomeric mutant derived from the tetrameric Discosoma sp. red fluorescent protein DsRed (1). DsRed-Monomer contains forty-five amino acid substitutions. When DsRed-Monomer is expressed in mammalian cell cultures, red fluorescent cells can be detected by either fluorescence microscopy or flow cytometry 12–16 hours after transfection or 24–48 hours after infection (DsRed-Monomer excitation and emission maxima = 557 nm and 592 nm, respectively. The DsRed-Monomer coding sequence is human codon-optimized for high expression in mammalian cells (2) The MCS in pRetroQ-DsRed Monomer-N1 lies between the immediate early promoter of CMV (PCMV IE) and the DsRed-Monomer coding sequences. Genes cloned into the MCS are expressed as fusions to the N-terminus of DsRed Monomer when they are in the same reading frame as DsRed Monomer and there are no intervening stop codons. The RetroQ retroviral vector backbone incorporates several unique features. This vector contains a puromycin resistance cassette (Puror) driven by the PGK promoter for selection of positively-infected cells (2).The hybrid 5’ long terminal repeat (LTR) consists of the CMV type I enhancer and the murine sarcoma virus (MSV) promoter. This vector demonstrates high levels of transcription in HEK 293-based packaging cell lines due, in part, to the presence of adenoviral E1A (3–6) in these cells. The self-inactivating feature of the vector is provided by a deletion in the 3’ LTR enhancer region (U3). During reverse transcription of the retroviral RNA, the inactivated 3’ LTR is copied and replaces the 5’ LTR CMV enhancer sequences. This can reduce the phenomenon known as promoter interference (7) and allow more efficient expression.
Additionally, the viral genomic transcript contains the necessary viral RNA processing elements, including the LTRs, packaging signal (ψ+), and tRNA primer-binding site. pRetroQDsRed Monomer-N1 contains a bacterial origin of replication, an E. coli Ampr gene for propagation and selection in bacteria, and an SV40 origin for replication in mammalian cells expressing the SV40 T antigen.
pRetroQ-DsRed Monomer-N1 is designed to efficiently deliver and express fusions to the N terminus of DsRed-Monomer into primary cells or cells that are difficult to transfect. Fusions to the N terminus of DsRed-Monomer retain the fluorescent properties of the native protein, allowing the in vivo localization of the fusion protein. The target gene should be cloned into pDsRed Monomer-N1 so that it is in frame with the DsRed-Monomer coding sequences, with no intervening in-frame stop codons. The inserted gene should include the initiating ATG codon. The recombinant DsRed Monomer vector can be infected or transfected into mammalian cells. If required, stable transformants can be selected using puromycin. pRetroQ DsRed Monomer-N1 can also be used simply to express DsRed Monomer in a cell line of interest (e.g., as an infection marker). Once pRetroQ-DsRed Monomer-N1 is transfected into a packaging cell line (such as the RetroPack PT67 Cell line, AmphoPack-293, EcoPack2-293, Pantropic Expression System , or Retro-X Universal Packaging System, RNA from the vector is packaged into non-infectious, replication-incompetent retroviral particles, since pRetroQ-DsRed Monomer-N1 lacks the structural genes (gag, pol, and env) necessary for particle formation and replication. These genes, however, are stably integrated as part of the packaging cell genome. Once a high-titer supernatant is produced, these retroviral particles can infect target cells and transmit the gene of interest but cannot replicate within these cells due to the absence of viral structural genes. The separate introduction and integration of the structural genes into the packaging cell line minimizes the chances of producing replicationcompetent virus due to recombination events during cell proliferation.
Propagation in E. coli
Suitable host strains: DH5α, Fusion Blue, and other general purpose strains.
Selectable marker: plasmid confers resistance to ampicillin (100 μg/ml) in E. coli hosts.
E. coli replication origin: ColE1
Copy number: low
pRetroQ-DsRed-Monomer-N1质粒序列载体序列
LOCUS pRetroQ-DsRed-Monomer-N1 7571 bp DNA SYN
DEFINITION pRetroQ-DsRed-Monomer-N1
ACCESSION
KEYWORDS
SOURCE
ORGANISM other sequences; artificial sequences; vectors.
FEATURES Location/Qualifiers
source 1..7571
/organism="pRetroQ-DsRed-Monomer-N1"
/mol_type="other DNA"
promoter 2..506
/label="CMV_immearly_promoter"
misc_feature 6..688
/label="5_LTR"
misc_feature 9..296
/label="CAG_enhancer"
misc_feature 463..483
/label="CMV_fwd_primer"
misc_feature 758..1567
/label="psi_plus_pack"
misc_feature 1154..1561
/label="gag"
misc_feature 1497..1519
/label="MSCV_primer"
misc_feature 1497..1519
/label="pLXSN_5_primer"
misc_feature 1535..1551
/label="pBABE_5_primer"
promoter 1591..2143
/label="CMV_immearly_promoter"
misc_feature 1646..1933
/label="CAG_enhancer"
misc_feature 2100..2120
/label="CMV_fwd_primer"
promoter 2101..2170
/label="CMV_promoter"
CDS 2260..2937
/label="ORF frame 1"
/translation="MDNTEDVIKEFMQFKVRMEGSVNGHYFEIEGEGEGKPYEGTQTA
KLQVTKGGPLPFAWDILSPQFQYGSKAYVKHPADIPDYMKLSFPEGFTWERSMNFEDG
GVVEVQQDSSLQDGTFIYKVKFKGVNFPADGPVMQKKTAGWEPSTEKLYPQDGVLKGE
ISHALKLKDGGHYTCDFKTVYKAKKPVQLPGNHYVDSKLDITNHNEDYTVVEQYEHAE
ARHSGSQ*"
gene 2263..2934
/label="DsRed_Monomer"
/gene="DsRed_Monomer"
/translation="MDNTEDVIKEFMQFKVRMEGSVNGHYFEIEGEGEGKPYEGTQTA
KLQVTKGGPLPFAWDILSPQFQYGSKAYVKHPADIPDYMKLSFPEGFTWERSMNFEDG
GVVEVQQDSSLQDGTFIYKVKFKGVNFPADGPVMQKKTAGWEPSTEKLYPQDGVLKGE
ISHALKLKDGGHYTCDFKTVYKAKKPVQLPGNHYVDSKLDITNHNEDYTVVEQYEHAE
ARHSGSQ*"
misc_feature complement(2440..2460)
/label="dsRed1_N_primer"
/translation="MDNTEDVIKEFMQFKVRMEGSVNGHYFEIEGEGEGKPYEGTQTA
KLQVTKGGPLPFAWDILSPQFQYGSKAYVKHPADIPDYMKLSFPEGFTWERSMNFEDG
GVVEVQQDSSLQDGTFIYKVKFKGVNFPADGPVMQKKTAGWEPSTEKLYPQDGVLKGE
ISHALKLKDGGHYTCDFKTVYKAKKPVQLPGNHYVDSKLDITNHNEDYTVVEQYEHAE
ARHSGSQ*"
misc_feature complement(3001..3024)
/label="MSCV_rev_primer"
/translation="MDNTEDVIKEFMQFKVRMEGSVNGHYFEIEGEGEGKPYEGTQTA
KLQVTKGGPLPFAWDILSPQFQYGSKAYVKHPADIPDYMKLSFPEGFTWERSMNFEDG
GVVEVQQDSSLQDGTFIYKVKFKGVNFPADGPVMQKKTAGWEPSTEKLYPQDGVLKGE
ISHALKLKDGGHYTCDFKTVYKAKKPVQLPGNHYVDSKLDITNHNEDYTVVEQYEHAE
ARHSGSQ*"
CDS 3246..4085
/label="ORF frame 3"
/translation="MEAGRPLGQRPIAALLLRFLGSEAGKGWVRGRAQGRAQGRGGRP
KVLRRPGILHASKAHVCRAVLLFLISGPFDLQPKLTMTEYKPTVRLATRDDVPRAVRT
LAAAFADYPATRHTVDPDRHIERVTELQELFLTRVGLDIGKVWVADDGAAVAVWTTPE
SVEAGAVFAEIGPRMAELSGSRLAAQQQMEGLLAPHRPKEPAWFLATVGVSPDHQGKG
LGSAVVLPGVEAAERAGVPAFLETSAPRNLPFYERLGFTVTADVEVPEGPRTWCMTRK
PGA*"
misc_feature 3398..3417
/label="mPGK_F_primer"
/translation="MEAGRPLGQRPIAALLLRFLGSEAGKGWVRGRAQGRAQGRGGRP
KVLRRPGILHASKAHVCRAVLLFLISGPFDLQPKLTMTEYKPTVRLATRDDVPRAVRT
LAAAFADYPATRHTVDPDRHIERVTELQELFLTRVGLDIGKVWVADDGAAVAVWTTPE
SVEAGAVFAEIGPRMAELSGSRLAAQQQMEGLLAPHRPKEPAWFLATVGVSPDHQGKG
LGSAVVLPGVEAAERAGVPAFLETSAPRNLPFYERLGFTVTADVEVPEGPRTWCMTRK
PGA*"
CDS complement(3411..4136)
/label="ORF frame 1"
/translation="MGSCAPFGRALRVVGRASGTGLAGHAPGARSFGHLDVGGDGEAE
PLVEGEVAGRGGLQEGGHPGALGRLHSGEHDGAAQTLALVVGRDADGGQEPRGLLGPV
RRQEAFHLLLRGQPGTAQLGHARADLGEHRPRFDALRRGPDRHRGAVVRDPHLADVEP
DAREEEFLQLGDPLDVAVRIDGVARGGVVGERGGEGAYGPGDVVAGGEAHRGLVLGHG
KLGLQVERPGDEEEENSAADVRF*"
gene 3486..4085
/label="puro (variant)"
/gene="puro (variant)"
/translation="MGSCAPFGRALRVVGRASGTGLAGHAPGARSFGHLDVGGDGEAE
PLVEGEVAGRGGLQEGGHPGALGRLHSGEHDGAAQTLALVVGRDADGGQEPRGLLGPV
RRQEAFHLLLRGQPGTAQLGHARADLGEHRPRFDALRRGPDRHRGAVVRDPHLADVEP
DAREEEFLQLGDPLDVAVRIDGVARGGVVGERGGEGAYGPGDVVAGGEAHRGLVLGHG
KLGLQVERPGDEEEENSAADVRF*"
misc_feature complement(4805..4827)
/label="pGEX_3_primer"
/translation="MGSCAPFGRALRVVGRASGTGLAGHAPGARSFGHLDVGGDGEAE
PLVEGEVAGRGGLQEGGHPGALGRLHSGEHDGAAQTLALVVGRDADGGQEPRGLLGPV
RRQEAFHLLLRGQPGTAQLGHARADLGEHRPRFDALRRGPDRHRGAVVRDPHLADVEP
DAREEEFLQLGDPLDVAVRIDGVARGGVVGERGGEGAYGPGDVVAGGEAHRGLVLGHG
KLGLQVERPGDEEEENSAADVRF*"
misc_feature complement(4963..4983)
/label="pBABE_3_primer"
/translation="MGSCAPFGRALRVVGRASGTGLAGHAPGARSFGHLDVGGDGEAE
PLVEGEVAGRGGLQEGGHPGALGRLHSGEHDGAAQTLALVVGRDADGGQEPRGLLGPV
RRQEAFHLLLRGQPGTAQLGHARADLGEHRPRFDALRRGPDRHRGAVVRDPHLADVEP
DAREEEFLQLGDPLDVAVRIDGVARGGVVGERGGEGAYGPGDVVAGGEAHRGLVLGHG
KLGLQVERPGDEEEENSAADVRF*"
misc_feature complement(4969..5184)
/label="SV40_enhancer"
/translation="MGSCAPFGRALRVVGRASGTGLAGHAPGARSFGHLDVGGDGEAE
PLVEGEVAGRGGLQEGGHPGALGRLHSGEHDGAAQTLALVVGRDADGGQEPRGLLGPV
RRQEAFHLLLRGQPGTAQLGHARADLGEHRPRFDALRRGPDRHRGAVVRDPHLADVEP
DAREEEFLQLGDPLDVAVRIDGVARGGVVGERGGEGAYGPGDVVAGGEAHRGLVLGHG
KLGLQVERPGDEEEENSAADVRF*"
promoter 4981..5249
/label="SV40_promoter"
/translation="MGSCAPFGRALRVVGRASGTGLAGHAPGARSFGHLDVGGDGEAE
PLVEGEVAGRGGLQEGGHPGALGRLHSGEHDGAAQTLALVVGRDADGGQEPRGLLGPV
RRQEAFHLLLRGQPGTAQLGHARADLGEHRPRFDALRRGPDRHRGAVVRDPHLADVEP
DAREEEFLQLGDPLDVAVRIDGVARGGVVGERGGEGAYGPGDVVAGGEAHRGLVLGHG
KLGLQVERPGDEEEENSAADVRF*"
rep_origin 5148..5225
/label="SV40_origin"
/translation="MGSCAPFGRALRVVGRASGTGLAGHAPGARSFGHLDVGGDGEAE
PLVEGEVAGRGGLQEGGHPGALGRLHSGEHDGAAQTLALVVGRDADGGQEPRGLLGPV
RRQEAFHLLLRGQPGTAQLGHARADLGEHRPRFDALRRGPDRHRGAVVRDPHLADVEP
DAREEEFLQLGDPLDVAVRIDGVARGGVVGERGGEGAYGPGDVVAGGEAHRGLVLGHG
KLGLQVERPGDEEEENSAADVRF*"
misc_feature 5210..5229
/label="SV40pro_F_primer"
/translation="MGSCAPFGRALRVVGRASGTGLAGHAPGARSFGHLDVGGDGEAE
PLVEGEVAGRGGLQEGGHPGALGRLHSGEHDGAAQTLALVVGRDADGGQEPRGLLGPV
RRQEAFHLLLRGQPGTAQLGHARADLGEHRPRFDALRRGPDRHRGAVVRDPHLADVEP
DAREEEFLQLGDPLDVAVRIDGVARGGVVGERGGEGAYGPGDVVAGGEAHRGLVLGHG
KLGLQVERPGDEEEENSAADVRF*"
gene complement(6349..7209)
/label="Ampicillin"
/gene="Ampicillin"
/translation="MGSCAPFGRALRVVGRASGTGLAGHAPGARSFGHLDVGGDGEAE
PLVEGEVAGRGGLQEGGHPGALGRLHSGEHDGAAQTLALVVGRDADGGQEPRGLLGPV
RRQEAFHLLLRGQPGTAQLGHARADLGEHRPRFDALRRGPDRHRGAVVRDPHLADVEP
DAREEEFLQLGDPLDVAVRIDGVARGGVVGERGGEGAYGPGDVVAGGEAHRGLVLGHG
KLGLQVERPGDEEEENSAADVRF*"
CDS complement(6349..7209)
/label="ORF frame 3"
/translation="MSIQHFRVALIPFFAAFCLPVFAHPETLVKVKDAEDQLGARVGY
IELDLNSGKILESFRPEERFPMMSTFKVLLCGAVLSRVDAGQEQLGRRIHYSQNDLVE
YSPVTEKHLTDGMTVRELCSAAITMSDNTAANLLLTTIGGPKELTAFLHNMGDHVTRL
DRWEPELNEAIPNDERDTTMPAAMATTLRKLLTGELLTLASRQQLIDWMEADKVAGPL
LRSALPAGWFIADKSGAGERGSRGIIAALGPDGKPSRIVVIYTTGSQATMDERNRQIA
EIGASLIKHW*"
promoter complement(7251..7279)
/label="AmpR_promoter"
/translation="MSIQHFRVALIPFFAAFCLPVFAHPETLVKVKDAEDQLGARVGY
IELDLNSGKILESFRPEERFPMMSTFKVLLCGAVLSRVDAGQEQLGRRIHYSQNDLVE
YSPVTEKHLTDGMTVRELCSAAITMSDNTAANLLLTTIGGPKELTAFLHNMGDHVTRL
DRWEPELNEAIPNDERDTTMPAAMATTLRKLLTGELLTLASRQQLIDWMEADKVAGPL
LRSALPAGWFIADKSGAGERGSRGIIAALGPDGKPSRIVVIYTTGSQATMDERNRQIA
EIGASLIKHW*"
promoter 7501..506
/label="CMV_immearly_promoter"
/translation="MSIQHFRVALIPFFAAFCLPVFAHPETLVKVKDAEDQLGARVGY
IELDLNSGKILESFRPEERFPMMSTFKVLLCGAVLSRVDAGQEQLGRRIHYSQNDLVE
YSPVTEKHLTDGMTVRELCSAAITMSDNTAANLLLTTIGGPKELTAFLHNMGDHVTRL
DRWEPELNEAIPNDERDTTMPAAMATTLRKLLTGELLTLASRQQLIDWMEADKVAGPL
LRSALPAGWFIADKSGAGERGSRGIIAALGPDGKPSRIVVIYTTGSQATMDERNRQIA
EIGASLIKHW*"
misc_feature 7565..688
/label="5_LTR"
/translation="MSIQHFRVALIPFFAAFCLPVFAHPETLVKVKDAEDQLGARVGY
IELDLNSGKILESFRPEERFPMMSTFKVLLCGAVLSRVDAGQEQLGRRIHYSQNDLVE
YSPVTEKHLTDGMTVRELCSAAITMSDNTAANLLLTTIGGPKELTAFLHNMGDHVTRL
DRWEPELNEAIPNDERDTTMPAAMATTLRKLLTGELLTLASRQQLIDWMEADKVAGPL
LRSALPAGWFIADKSGAGERGSRGIIAALGPDGKPSRIVVIYTTGSQATMDERNRQIA
EIGASLIKHW*"
ORIGIN
1 TCCGCGTTAC ATAACTTACG GTAAATGGCC CGCCTGGCTG ACCGCCCAAC GACCCCCGCC
61 CATTGACGTC AATAATGACG TATGTTCCCA TAGTAACGCC AATAGGGACT TTCCATTGAC
121 GTCAATGGGT GGAGTATTTA CGGTAAACTG CCCACTTGGC AGTACATCAA GTGTATCATA
181 TGCCAAGTAC GCCCCCTATT GACGTCAATG ACGGTAAATG GCCCGCCTGG CATTATGCCC
241 AGTACATGAC CTTATGGGAC TTTCCTACTT GGCAGTACAT CTACGTATTA GTCATCGCTA
301 TTACCATGGT GATGCGGTTT TGGCAGTACA TCAATGGGCG TGGATAGCGG TTTGACTCAC
361 GGGGATTTCC AAGTCTCCAC CCCATTGACG TCAATGGGAG TTTGTTTTGG CACCAAAATC
421 AACGGGACTT TCCAAAATGT CGTAACAACT CCGCCCCATT GACGCAAATG GGCGGTAGGC
481 GTGTACGGTG GGAGGTCTAT ATAAGCAGAG CTCAATAAAA GAGCCCACAA CCCCTCACTC
541 GGCGCGCCAG TCTTCCGATA GACTGCGTCG CCCGGGTACC CGTATTCCCA ATAAAGCCTC
601 TTGCTGTTTG CATCCGAATC GTGGTCTCGC TGTTCCTTGG GAGGGTCTCC TCTGAGTGAT
661 TGACTACCCA CGACGGGGGT CTTTCATTTG GGGGCTCGTC CGGGATTTGG AGACCCCTGC
721 CCAGGGACCA CCGACCCACC ACCGGGAGGT AAGCTGGCCA GCAACTTATC TGTGTCTGTC
781 CGATTGTCTA GTGTCTATGT TTGATGTTAT GCGCCTGCGT CTGTACTAGT TAGCTAACTA
841 GCTCTGTATC TGGCGGACCC GTGGTGGAAC TGACGAGTTC TGAACACCCG GCCGCAACCC
901 TGGGAGACGT CCCAGGGACT TTGGGGGCCG TTTTTGTGGC CCGACCTGAG GAAGGGAGTC
961 GATGTGGAAT CCGACCCCGT CAGGATATGT GGTTCTGGTA GGAGACGAGA ACCTAAAACA
1021 GTTCCCGCCT CCGTCTGAAT TTTTGCTTTC GGTTTGGAAC CGAAGCCGCG CGTCTTGTCT
1081 GCTGCAGCGC TGCAGCATCG TTCTGTGTTG TCTCTGTCTG ACTGTGTTTC TGTATTTGTC
1141 TGAAAATTAG GGCCAGACTG TTACCACTCC CTTAAGTTTG ACCTTAGGTC ACTGGAAAGA
1201 TGTCGAGCGG ATCGCTCACA ACCAGTCGGT AGATGTCAAG AAGAGACGTT GGGTTACCTT
1261 CTGCTCTGCA GAATGGCCAA CCTTTAACGT CGGATGGCCG CGAGACGGCA CCTTTAACCG
1321 AGACCTCATC ACCCAGGTTA AGATCAAGGT CTTTTCACCT GGCCCGCATG GACACCCAGA
1381 CCAGGTCCCC TACATCGTGA CCTGGGAAGC CTTGGCTTTT GACCCCCCTC CCTGGGTCAA
1441 GCCCTTTGTA CACCCTAAGC CTCCGCCTCC TCTTCCTCCA TCCGCCCCGT CTCTCCCCCT
1501 TGAACCTCCT CGTTCGACCC CGCCTCGATC CTCCCTTTAT CCAGCCCTCA CTCCTTCTCT
1561 AGGCGCCGGA ATTGAAGATC ATAGTTATTA ATAGTAATCA ATTACGGGGT CATTAGTTCA
1621 TAGCCCATAT ATGGAGTTCC GCGTTACATA ACTTACGGTA AATGGCCCGC CTGGCTGACC
1681 GCCCAACGAC CCCCGCCCAT TGACGTCAAT AATGACGTAT GTTCCCATAG TAACGCCAAT
1741 AGGGACTTTC CATTGACGTC AATGGGTGGA GTATTTACGG TAAACTGCCC ACTTGGCAGT
1801 ACATCAAGTG TATCATATGC CAAGTACGCC CCCTATTGAC GTCAATGACG GTAAATGGCC
1861 CGCCTGGCAT TATGCCCAGT ACATGACCTT ATGGGACTTT CCTACTTGGC AGTACATCTA
1921 CGTATTAGTC ATCGCTATTA CCATGGTGAT GCGGTTTTGG CAGTACATCA ATGGGCGTGG
1981 ATAGCGGTTT GACTCACGGG GATTTCCAAG TCTCCACCCC ATTGACGTCA ATGGGAGTTT
2041 GTTTTGGCAC CAAAATCAAC GGGACTTTCC AAAATGTCGT AACAACTCCG CCCCATTGAC
2101 GCAAATGGGC GGTAGGCGTG TACGGTGGGA GGTCTATATA AGCAGAGCTG GTTTAGTGAA
2161 CCGTCAGATC CGCTAGCGCT ACCGGACTCA GATCTCGAGC TCAAGCTTCG AATTCTGCAG
2221 TCGACGGTAC CGCGGGCCCG GGATCCACCG GTCGCCACCA TGGACAACAC CGAGGACGTC
2281 ATCAAGGAGT TCATGCAGTT CAAGGTGCGC ATGGAGGGCT CCGTGAACGG CCACTACTTC
2341 GAGATCGAGG GCGAGGGCGA GGGCAAGCCC TACGAGGGCA CCCAGACCGC CAAGCTGCAG
2401 GTGACCAAGG GCGGCCCCCT GCCCTTCGCC TGGGACATCC TGTCCCCCCA GTTCCAGTAC
2461 GGCTCCAAGG CCTACGTGAA GCACCCCGCC GACATCCCCG ACTACATGAA GCTGTCCTTC
2521 CCCGAGGGCT TCACCTGGGA GCGCTCCATG AACTTCGAGG ACGGCGGCGT GGTGGAGGTG
2581 CAGCAGGACT CCTCCCTGCA GGACGGCACC TTCATCTACA AGGTGAAGTT CAAGGGCGTG
2641 AACTTCCCCG CCGACGGCCC CGTAATGCAG AAGAAGACTG CCGGCTGGGA GCCCTCCACC
2701 GAGAAGCTGT ACCCCCAGGA CGGCGTGCTG AAGGGCGAGA TCTCCCACGC CCTGAAGCTG
2761 AAGGACGGCG GCCACTACAC CTGCGACTTC AAGACCGTGT ACAAGGCCAA GAAGCCCGTG
2821 CAGCTGCCCG GCAACCACTA CGTGGACTCC AAGCTGGACA TCACCAACCA CAACGAGGAC
2881 TACACCGTGG TGGAGCAGTA CGAGCACGCC GAGGCCCGCC ACTCCGGCTC CCAGTAGAGC
2941 GGCCGCGACT CTAGATAATT CTACCGGGTA GGGGAGGCGC TTTTCCCAAG GCAGTCTGGA
3001 GCATGCGCTT TAGCAGCCCC GCTGGGCACT TGGCGCTACA CAAGTGGCCT CTGGCCTCGC
3061 ACACATTCCA CATCCACCGG TAGGCGCCAA CCGGCTCCGT TCTTTGGTGG CCCCTTCGCG
3121 CCACCTTCTA CTCCTCCCCT AGTCAGGAAG TTCCCCCCCG CCCCGCAGCT CGCGTCGTGC
3181 AGGACGTGAC AAATGGAAGT AGCACGTCTC ACTAGTCTCG TGCAGATGGA CAGCACCGCT
3241 GAGCAATGGA AGCGGGTAGG CCTTTGGGGC AGCGGCCAAT AGCAGCTTTG CTCCTTCGCT
3301 TTCTGGGCTC AGAGGCTGGG AAGGGGTGGG TCCGGGGGCG GGCTCAGGGG CGGGCTCAGG
3361 GGCGGGGCGG GCGCCCGAAG GTCCTCCGGA GGCCCGGCAT TCTGCACGCT TCAAAAGCGC
3421 ACGTCTGCCG CGCTGTTCTC CTCTTCCTCA TCTCCGGGCC TTTCGACCTG CAGCCCAAGC
3481 TTACCATGAC CGAGTACAAG CCCACGGTGC GCCTCGCCAC CCGCGACGAC GTCCCCAGGG
3541 CCGTACGCAC CCTCGCCGCC GCGTTCGCCG ACTACCCCGC CACGCGCCAC ACCGTCGATC
3601 CGGACCGCCA CATCGAGCGG GTCACCGAGC TGCAAGAACT CTTCCTCACG CGCGTCGGGC
3661 TCGACATCGG CAAGGTGTGG GTCGCGGACG ACGGCGCCGC GGTGGCGGTC TGGACCACGC
3721 CGGAGAGCGT CGAAGCGGGG GCGGTGTTCG CCGAGATCGG CCCGCGCATG GCCGAGTTGA
3781 GCGGTTCCCG GCTGGCCGCG CAGCAACAGA TGGAAGGCCT CCTGGCGCCG CACCGGCCCA
3841 AGGAGCCCGC GTGGTTCCTG GCCACCGTCG GCGTCTCGCC CGACCACCAG GGCAAGGGTC
3901 TGGGCAGCGC CGTCGTGCTC CCCGGAGTGG AGGCGGCCGA GCGCGCCGGG GTGCCCGCCT
3961 TCCTGGAGAC CTCCGCGCCC CGCAACCTCC CCTTCTACGA GCGGCTCGGC TTCACCGTCA
4021 CCGCCGACGT CGAGGTGCCC GAAGGACCGC GCACCTGGTG CATGACCCGC AAGCCCGGTG
4081 CCTGACGCCC GCCCCACGAC CCGCAGCGCC CGACCGAAAG GAGCGCACGA CCCCATGCAT
4141 CGGCGTCTCG ATCGAGATAT CAGTGGTCCA GGCTCTAGTT TTGACTCAAC AATATCACCA
4201 GCTGAAGCCT ATAGAGTACG AGCCATAGAT AAAATAAAAG ATTTTATTTA GTCTCCAGAA
4261 AAAGGGGGGA ATGAAAGACC CCACCTGTAG GTTTGGCAAG CTACTAGCTT AAGTAACGCC
4321 ATTTTGCAAG GCATGGAAAA ATACATAACT GAGAATAGAG AAGTTCAGAT CAAGGTCAGG
4381 AACAGATGGA ACAGGGTCGA TCGACCCTAG AGAACCATCA GATGTTTCCA GGGTGCCCCA
4441 AGGACCTGAA ATGACCCTGT GCCTTATTTG AACTAACCAA TCAGTTCGCT TCTCGCTTCT
4501 GTTCGCGCGC TTCTGCTCCC CGAGCTCAAT AAAAGAGCCC ACAACCCCTC ACTCGGGGCG
4561 CCAGTCCTCC GATTGACTGA GTCGCCCGGG TACCCGTGTA TCCAATAAAC CCTCTTGCAG
4621 TTGCATCCGA CTTGTGGTCT CGCTGTTCCT TGGGAGGGTC TCCTCTGAGT GATTGACTAC
4681 CCGTCAGCGG GGGTCTTTCA TTTGGGGGCT CGTCCGGGAT CGGGAGACCC CTGCCCAGGG
4741 ACCACCGACC CACCACCGGG AGGTAAGCTG GCTGCCTCGC GCGTTTCGGT GATGACGGTG
4801 AAAACCTCTG ACACATGCAG CTCCCGGAGA CGGTCACAGC TTGTCTGTAA GCGGATGCCG
4861 GGAGCAGACA AGCCCGTCAG GGCGCGTCAG CGGGTGTTGG CGGGTGTCGG GGCGCAGCCA
4921 TGACCCAGTC ACGTAGCGAT AGCGGAGTGT AGATCCGGCT GTGGAATGTG TGTCAGTTAG
4981 GGTGTGGAAA GTCCCCAGGC TCCCCAGCAG GCAGAAGTAT GCAAAGCATG CATCTCAATT
5041 AGTCAGCAAC CAGGTGTGGA AAGTCCCCAG GCTCCCCAGC AGGCAGAAGT ATGCAAAGCA
5101 TGCATCTCAA TTAGTCAGCA ACCATAGTCC CGCCCCTAAC TCCGCCCATC CCGCCCCTAA
5161 CTCCGCCCAG TTCCGCCCAT TCTCCGCCCC ATGGCTGACT AATTTTTTTT ATTTATGCAG
5221 AGGCCGAGGC CGCCTCGGCC TCTGAGCTAT TCCAGAAGTA GTGAGGAGGC TTTTTTGGAG
5281 GCCTAGGCTT TTGCAAAAAG CTTACTGGCT TAACTATGCG GCATCAGAGC AGATTGTACT
5341 GAGAGTGCAC CATATGCGGT GTGAAATACC GCACAGATGC GTAAGGAGAA AATACCGCAT
5401 CAGGCGCTCT TCCGCTTCCT CGCTCACTGA CTCGCTGCGC TCGGTCGTTC GGCTGCGGCG
5461 AGCGGTATCA GCTCACTCAA AGGCGGTAAT ACGGTTATCC ACAGAATCAG GGGATAACGC
5521 AGGAAAGAAC ATGTGAGCAA AAGGCCAGCA AAAGGCCAGG AACCGTAAAA AGGCCGCGTT
5581 GCTGGCGTTT TTCCATAGGC TCCGCCCCCC TGACGAGCAT CACAAAAATC GACGCTCAAG
5641 TCAGAGGTGG CGAAACCCGA CAGGACTATA AAGATACCAG GCGTTTCCCC CTGGAAGCTC
5701 CCTCGTGCGC TCTCCTGTTC CGACCCTGCC GCTTACCGGA TACCTGTCCG CCTTTCTCCC
5761 TTCGGGAAGC GTGGCGCTTT CTCATAGCTC ACGCTGTAGG TATCTCAGTT CGGTGTAGGT
5821 CGTTCGCTCC AAGCTGGGCT GTGTGCACGA ACCCCCCGTT CAGCCCGACC GCTGCGCCTT
5881 ATCCGGTAAC TATCGTCTTG AGTCCAACCC GGTAAGACAC GACTTATCGC CACTGGCAGC
5941 AGCCACTGGT AACAGGATTA GCAGAGCGAG GTATGTAGGC GGTGCTACAG AGTTCTTGAA
6001 GTGGTGGCCT AACTACGGCT ACACTAGAAG GACAGTATTT GGTATCTGCG CTCTGCTGAA
6061 GCCAGTTACC TTCGGAAAAA GAGTTGGTAG CTCTTGATCC GGCAAACAAA CCACCGCTGG
6121 TAGCGGTGGT TTTTTTGTTT GCAAGCAGCA GATTACGCGC AGAAAAAAAG GATCTCAAGA
6181 AGATCCTTTG ATCTTTTCTA CGGGGTCTGA CGCTCAGTGG AACGAAAACT CACGTTAAGG
6241 GATTTTGGTC ATGAGATTAT CAAAAAGGAT CTTCACCTAG ATCCTTTTAA ATTAAAAATG
6301 AAGTTTTAAA TCAATCTAAA GTATATATGA GTAAACTTGG TCTGACAGTT ACCAATGCTT
6361 AATCAGTGAG GCACCTATCT CAGCGATCTG TCTATTTCGT TCATCCATAG TTGCCTGACT
6421 CCCCGTCGTG TAGATAACTA CGATACGGGA GGGCTTACCA TCTGGCCCCA GTGCTGCAAT
6481 GATACCGCGA GACCCACGCT CACCGGCTCC AGATTTATCA GCAATAAACC AGCCAGCCGG
6541 AAGGGCCGAG CGCAGAAGTG GTCCTGCAAC TTTATCCGCC TCCATCCAGT CTATTAATTG
6601 TTGCCGGGAA GCTAGAGTAA GTAGTTCGCC AGTTAATAGT TTGCGCAACG TTGTTGCCAT
6661 TGCTGCAGGC ATCGTGGTGT CACGCTCGTC GTTTGGTATG GCTTCATTCA GCTCCGGTTC
6721 CCAACGATCA AGGCGAGTTA CATGATCCCC CATGTTGTGC AAAAAAGCGG TTAGCTCCTT
6781 CGGTCCTCCG ATCGTTGTCA GAAGTAAGTT GGCCGCAGTG TTATCACTCA TGGTTATGGC
6841 AGCACTGCAT AATTCTCTTA CTGTCATGCC ATCCGTAAGA TGCTTTTCTG TGACTGGTGA
6901 GTACTCAACC AAGTCATTCT GAGAATAGTG TATGCGGCGA CCGAGTTGCT CTTGCCCGGC
6961 GTCAACACGG GATAATACCG CGCCACATAG CAGAACTTTA AAAGTGCTCA TCATTGGAAA
7021 ACGTTCTTCG GGGCGAAAAC TCTCAAGGAT CTTACCGCTG TTGAGATCCA GTTCGATGTA
7081 ACCCACTCGT GCACCCAACT GATCTTCAGC ATCTTTTACT TTCACCAGCG TTTCTGGGTG
7141 AGCAAAAACA GGAAGGCAAA ATGCCGCAAA AAAGGGAATA AGGGCGACAC GGAAATGTTG
7201 AATACTCATA CTCTTCCTTT TTCAATATTA TTGAAGCATT TATCAGGGTT ATTGTCTCAT
7261 GAGCGGATAC ATATTTGAAT GTATTTAGAA AAATAAACAA ATAGGGGTTC CGCGCACATT
7321 TCCCCGAAAA GTGCCACCTG ACGTCTAAGA AACCATTATT ATCATGACAT TAACCTATAA
7381 AAATAGGCGT ATCACGAGGC CCTTTCGTCT TCAAGAATTA GCTTGGCCAT TGCATACGTT
7441 GTATCCATAT CATAATATGT ACATTTATAT TGGCTCATGT CCAACATTAC CGCCATGTTG
7501 ACATTGATTA TTGACTAGTT ATTAATAGTA ATCAATTACG GGGTCATTAG TTCATAGCCC
7561 ATATATGGAG T
//