| 质粒名称: | mCherry-Peroxisomes-2 |
|---|---|
| 出品公司: | Addgene |
| 目录编号: | 54520 |
| 存储实验室: | Michael Davidson |
|
载体骨架基本信息 |
|
| 载体名称: | mCherry |
| 空载体大小: | 4750 bp |
| 载体类型: | 哺乳细胞表达 |
| 筛选标记: | 新霉素(G418) |
| 载体抗性: | 卡那霉素,50μg/mL |
| 生长温度: | 37 ℃ |
| 宿主菌: | DH5a |
| 拷贝数: | 高拷贝 |
|
插入基因信息 |
|
| 插入基因名称: | mCherry-Peroxisomes |
| 标签/融合蛋白: | 过氧化物酶体靶向信号1(PTS1;Ser-Lys-Leu)(插入物上的C末端) |
|
文献引用方法 |
|
| 材料和方法部分: | mCherry-Peroxisomes-2 was a gift from Michael Davidson (Addgene plasmid # 54520 ; http://n2t.net/addgene:54520 ; RRID:Addgene_54520) |
mCherry-Peroxisomes-2
价格:元
规格: 2ug质粒(仅用于转化提取) 转染级去内毒素液体即用质粒100μg 转染级去内毒素液体即用质粒500μg 转染级去内毒素液体即用质粒1mg 更大规格-质粒构建-质粒定制-请联系我们
联系方式:I47-825O-882O
买家导航

mCherry-Peroxisomes-2载体质粒基本信息
mCherry-Peroxisomes-2质粒图谱载体图谱和mCherry-Peroxisomes-2载体序列质粒序列多克隆位点信息
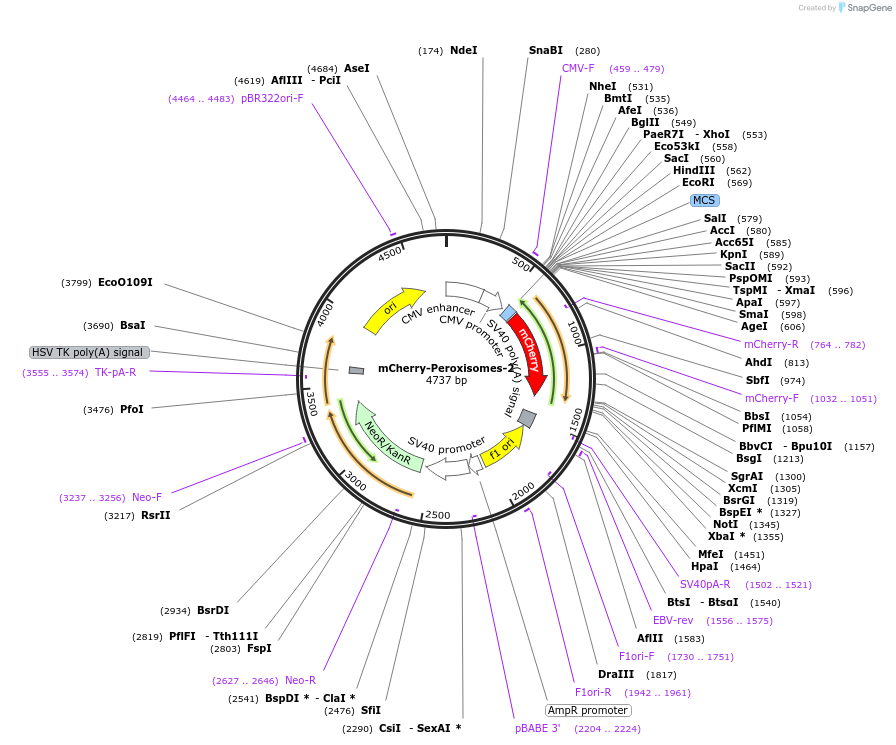
mCherry-Peroxisomes-2质粒载体简介
mCherry-Peroxisomes-2质粒序列载体序列
LOCUS mCherry-Peroxisomes-2 4737 bp DNA circular SYN 15-DEC-2025
DEFINITION Localization: Peroxisomes, Excitation: 587, Emission: 610.
ACCESSION .
VERSION .
KEYWORDS .
SOURCE synthetic DNA construct
ORGANISM synthetic DNA construct
REFERENCE 1 (bases 1 to 4737)
AUTHORS Davidson Lab
TITLE Michael Davidson Empty Backbones
JOURNAL unpublished
REFERENCE 2 (bases 1 to 4737)
AUTHORS .
TITLE Direct Submission
JOURNAL Exported Dec 15, 2025 from SnapGene Server 8.0.1
https://www.biofeng.com
COMMENT SGRef: number: 1; type: "Journal Article"; journalName:
"unpublished"
COMMENT Sequence Label: mCherry-Peroxisomes-2
FEATURES Location/Qualifiers
source 1..4737
/mol_type="other DNA"
/organism="synthetic DNA construct"
enhancer 1..304
/label=CMV enhancer
/note="human cytomegalovirus immediate early enhancer"
promoter 305..508
/label=CMV promoter
/note="human cytomegalovirus (CMV) immediate early
promoter"
primer_bind 459..479
/label=CMV-F
/note="Human CMV immediate early promoter, forward primer"
misc_feature 531..611
/label=MCS
/note="multiple cloning site"
regulatory 613..622
/label=Kozak sequence
/note="vertebrate consensus sequence for strong initiation
of translation (Kozak, 1987)"
/regulatory_class="other"
CDS 619..1326
/codon_start=1
/product="monomeric derivative of DsRed fluorescent protein
(Shaner et al., 2004)"
/label=mCherry
/note="mammalian codon-optimized"
/translation="MVSKGEEDNMAIIKEFMRFKVHMEGSVNGHEFEIEGEGEGRPYEG
TQTAKLKVTKGGPLPFAWDILSPQFMYGSKAYVKHPADIPDYLKLSFPEGFKWERVMNF
EDGGVVTVTQDSSLQDGEFIYKVKLRGTNFPSDGPVMQKKTMGWEASSERMYPEDGALK
GEIKQRLKLKDGGHYDAEVKTTYKAKKPVQLPGAYNVNIKLDITSHNEDYTIVEQYERA
EGRHSTGGMDELYK"
primer_bind complement(764..782)
/label=mCherry-R
/note="mCherry, reverse primer"
primer_bind 1032..1051
/label=mCherry-F
/note="mCherry, forward primer"
polyA_signal 1465..1586
/label=SV40 poly(A) signal
/note="SV40 polyadenylation signal"
primer_bind complement(1502..1521)
/label=SV40pA-R
/note="SV40 polyA, reverse primer"
primer_bind 1556..1575
/label=EBV-rev
/note="SV40 polyA terminator, reverse primer"
rep_origin complement(1593..2048)
/direction=LEFT
/label=f1 ori
/note="f1 bacteriophage origin of replication; arrow
indicates direction of (+) strand synthesis"
primer_bind complement(1730..1751)
/label=F1ori-F
/note="F1 origin, forward primer"
primer_bind 1942..1961
/label=F1ori-R
/note="F1 origin, reverse primer"
promoter 2075..2179
/gene="bla"
/label=AmpR promoter
promoter 2181..2538
/label=SV40 promoter
/note="SV40 enhancer and early promoter"
enhancer 2181..2372
/label=SV40 enhancer
/note="enhancer for the SV40 early promoter (Herr, 1993)"
primer_bind complement(2204..2224)
/label=pBABE 3'
/note="SV40 enhancer, reverse primer for pBABE vectors"
rep_origin 2389..2524
/label=SV40 ori
/note="SV40 origin of replication"
CDS 2573..3367
/codon_start=1
/product="aminoglycoside phosphotransferase"
/label=NeoR/KanR
/note="confers resistance to neomycin, kanamycin, and G418
(Geneticin(R))"
/translation="MIEQDGLHAGSPAAWVERLFGYDWAQQTIGCSDAAVFRLSAQGRP
VLFVKTDLSGALNELQDEAARLSWLATTGVPCAAVLDVVTEAGRDWLLLGEVPGQDLLS
SHLAPAEKVSIMADAMRRLHTLDPATCPFDHQAKHRIERARTRMEAGLVDQDDLDEEHQ
GLAPAELFARLKASMPDGEDLVVTHGDACLPNIMVENGRFSGFIDCGRLGVADRYQDIA
LATRDIAEELGGEWADRFLVLYGIAAPDSQRIAFYRLLDEFF"
CDS 2573..3367
/codon_start=1
/gene="aph(3')-II (or nptII)"
/product="aminoglycoside phosphotransferase from Tn5"
/label=NeoR/KanR
/note="confers resistance to neomycin, kanamycin, and G418
(Geneticin(R))"
/translation="MIEQDGLHAGSPAAWVERLFGYDWAQQTIGCSDAAVFRLSAQGRP
VLFVKTDLSGALNELQDEAARLSWLATTGVPCAAVLDVVTEAGRDWLLLGEVPGQDLLS
SHLAPAEKVSIMADAMRRLHTLDPATCPFDHQAKHRIERARTRMEAGLVDQDDLDEEHQ
GLAPAELFARLKASMPDGEDLVVTHGDACLPNIMVENGRFSGFIDCGRLGVADRYQDIA
LATRDIAEELGGEWADRFLVLYGIAAPDSQRIAFYRLLDEFF"
primer_bind complement(2627..2646)
/label=Neo-R
/note="Neomycin resistance gene, reverse primer"
primer_bind 3237..3256
/label=Neo-F
/note="Neomycin resistance gene, forward primer"
primer_bind complement(3555..3574)
/label=TK-pA-R
/note="Thymidine kinase polyA, reverse primer"
polyA_signal 3599..3646
/label=HSV TK poly(A) signal
/note="herpes simplex virus thymidine kinase
polyadenylation signal (Cole and Stacy, 1985)"
rep_origin 3975..4563
/direction=RIGHT
/label=ori
/note="high-copy-number ColE1/pMB1/pBR322/pUC origin of
replication"
primer_bind 4464..4483
/label=pBR322ori-F
/note="pBR322 origin, forward primer"
ORIGIN
1 cgttacataa cttacggtaa atggcccgcc tggctgaccg cccaacgacc cccgcccatt
61 gacgtcaata atgacgtatg ttcccatagt aacgccaata gggactttcc attgacgtca
121 atgggtggag tatttacggt aaactgccca cttggcagta catcaagtgt atcatatgcc
181 aagtacgccc cctattgacg tcaatgacgg taaatggccc gcctggcatt atgcccagta
241 catgacctta tgggactttc ctacttggca gtacatctac gtattagtca tcgctattac
301 catggtgatg cggttttggc agtacatcaa tgggcgtgga tagcggtttg actcacgggg
361 atttccaagt ctccacccca ttgacgtcaa tgggagtttg ttttggcacc aaaatcaacg
421 ggactttcca aaatgtcgta acaactccgc cccattgacg caaatgggcg gtaggcgtgt
481 acggtgggag gtctatataa gcagagctgg tttagtgaac cgtcagatcc gctagcgcta
541 ccggactcag atctcgagct caagcttcga attctgcagt cgacggtacc gcgggcccgg
601 gatccaccgg tcgccaccat ggtgagcaag ggcgaggagg ataacatggc catcatcaag
661 gagttcatgc gcttcaaggt gcacatggag ggctccgtga acggccacga gttcgagatc
721 gagggcgagg gcgagggccg cccctacgag ggcacccaga ccgccaagct gaaggtgacc
781 aagggtggcc ccctgccctt cgcctgggac atcctgtccc ctcagttcat gtacggctcc
841 aaggcctacg tgaagcaccc cgccgacatc cccgactact tgaagctgtc cttccccgag
901 ggcttcaagt gggagcgcgt gatgaacttc gaggacggcg gcgtggtgac cgtgacccag
961 gactcctccc tgcaggacgg cgagttcatc tacaaggtga agctgcgcgg caccaacttc
1021 ccctccgacg gccccgtaat gcagaagaag accatgggct gggaggcctc ctccgagcgg
1081 atgtaccccg aggacggcgc cctgaagggc gagatcaagc agaggctgaa gctgaaggac
1141 ggcggccact acgacgctga ggtcaagacc acctacaagg ccaagaagcc cgtgcagctg
1201 cccggcgcct acaacgtcaa catcaagttg gacatcacct cccacaacga ggactacacc
1261 atcgtggaac agtacgaacg cgccgagggc cgccactcca ccggcggcat ggacgagctg
1321 tacaagtccg gatccaagct gtagcggccg cgactctaga tcataatcag ccataccaca
1381 tttgtagagg ttttacttgc tttaaaaaac ctcccacacc tccccctgaa cctgaaacat
1441 aaaatgaatg caattgttgt tgttaacttg tttattgcag cttataatgg ttacaaataa
1501 agcaatagca tcacaaattt cacaaataaa gcattttttt cactgcattc tagttgtggt
1561 ttgtccaaac tcatcaatgt atcttaaggc gtaaattgta agcgttaata ttttgttaaa
1621 attcgcgtta aatttttgtt aaatcagctc attttttaac caataggccg aaatcggcaa
1681 aatcccttat aaatcaaaag aatagaccga gatagggttg agtgttgttc cagtttggaa
1741 caagagtcca ctattaaaga acgtggactc caacgtcaaa gggcgaaaaa ccgtctatca
1801 gggcgatggc ccactacgtg aaccatcacc ctaatcaagt tttttggggt cgaggtgccg
1861 taaagcacta aatcggaacc ctaaagggag cccccgattt agagcttgac ggggaaagcc
1921 ggcgaacgtg gcgagaaagg aagggaagaa agcgaaagga gcgggcgcta gggcgctggc
1981 aagtgtagcg gtcacgctgc gcgtaaccac cacacccgcc gcgcttaatg cgccgctaca
2041 gggcgcgtca ggtggcactt ttcggggaaa tgtgcgcgga acccctattt gtttattttt
2101 ctaaatacat tcaaatatgt atccgctcat gagacaataa ccctgataaa tgcttcaata
2161 atattgaaaa aggaagagtc ctgaggcgga aagaaccagc tgtggaatgt gtgtcagtta
2221 gggtgtggaa agtccccagg ctccccagca ggcagaagta tgcaaagcat gcatctcaat
2281 tagtcagcaa ccaggtgtgg aaagtcccca ggctccccag caggcagaag tatgcaaagc
2341 atgcatctca attagtcagc aaccatagtc ccgcccctaa ctccgcccat cccgccccta
2401 actccgccca gttccgccca ttctccgccc catggctgac taattttttt tatttatgca
2461 gaggccgagg ccgcctcggc ctctgagcta ttccagaagt agtgaggagg cttttttgga
2521 ggcctaggct tttgcaaaga tcgatcaaga gacaggatga ggatcgtttc gcatgattga
2581 acaagatgga ttgcacgcag gttctccggc cgcttgggtg gagaggctat tcggctatga
2641 ctgggcacaa cagacaatcg gctgctctga tgccgccgtg ttccggctgt cagcgcaggg
2701 gcgcccggtt ctttttgtca agaccgacct gtccggtgcc ctgaatgaac tgcaagacga
2761 ggcagcgcgg ctatcgtggc tggccacgac gggcgttcct tgcgcagctg tgctcgacgt
2821 tgtcactgaa gcgggaaggg actggctgct attgggcgaa gtgccggggc aggatctcct
2881 gtcatctcac cttgctcctg ccgagaaagt atccatcatg gctgatgcaa tgcggcggct
2941 gcatacgctt gatccggcta cctgcccatt cgaccaccaa gcgaaacatc gcatcgagcg
3001 agcacgtact cggatggaag ccggtcttgt cgatcaggat gatctggacg aagagcatca
3061 ggggctcgcg ccagccgaac tgttcgccag gctcaaggcg agcatgcccg acggcgagga
3121 tctcgtcgtg acccatggcg atgcctgctt gccgaatatc atggtggaaa atggccgctt
3181 ttctggattc atcgactgtg gccggctggg tgtggcggac cgctatcagg acatagcgtt
3241 ggctacccgt gatattgctg aagagcttgg cggcgaatgg gctgaccgct tcctcgtgct
3301 ttacggtatc gccgctcccg attcgcagcg catcgccttc tatcgccttc ttgacgagtt
3361 cttctgagcg ggactctggg gttcgaaatg accgaccaag cgacgcccaa cctgccatca
3421 cgagatttcg attccaccgc cgccttctat gaaaggttgg gcttcggaat cgttttccgg
3481 gacgccggct ggatgatcct ccagcgcggg gatctcatgc tggagttctt cgcccaccct
3541 agggggaggc taactgaaac acggaaggag acaataccgg aaggaacccg cgctatgacg
3601 gcaataaaaa gacagaataa aacgcacggt gttgggtcgt ttgttcataa acgcggggtt
3661 cggtcccagg gctggcactc tgtcgatacc ccaccgagac cccattgggg ccaatacgcc
3721 cgcgtttctt ccttttcccc accccacccc ccaagttcgg gtgaaggccc agggctcgca
3781 gccaacgtcg gggcggcagg ccctgccata gcctcaggtt actcatatat actttagatt
3841 gatttaaaac ttcattttta atttaaaagg atctaggtga agatcctttt tgataatctc
3901 atgaccaaaa tcccttaacg tgagttttcg ttccactgag cgtcagaccc cgtagaaaag
3961 atcaaaggat cttcttgaga tccttttttt ctgcgcgtaa tctgctgctt gcaaacaaaa
4021 aaaccaccgc taccagcggt ggtttgtttg ccggatcaag agctaccaac tctttttccg
4081 aaggtaactg gcttcagcag agcgcagata ccaaatactg ttcttctagt gtagccgtag
4141 ttaggccacc acttcaagaa ctctgtagca ccgcctacat acctcgctct gctaatcctg
4201 ttaccagtgg ctgctgccag tggcgataag tcgtgtctta ccgggttgga ctcaagacga
4261 tagttaccgg ataaggcgca gcggtcgggc tgaacggggg gttcgtgcac acagcccagc
4321 ttggagcgaa cgacctacac cgaactgaga tacctacagc gtgagctatg agaaagcgcc
4381 acgcttcccg aagggagaaa ggcggacagg tatccggtaa gcggcagggt cggaacagga
4441 gagcgcacga gggagcttcc agggggaaac gcctggtatc tttatagtcc tgtcgggttt
4501 cgccacctct gacttgagcg tcgatttttg tgatgctcgt caggggggcg gagcctatgg
4561 aaaaacgcca gcaacgcggc ctttttacgg ttcctggcct tttgctggcc ttttgctcac
4621 atgttctttc ctgcgttatc ccctgattct gtggataacc gtattaccgc catgcattag
4681 ttattaatag taatcaatta cggggtcatt agttcatagc ccatatatgg agttccg
//

