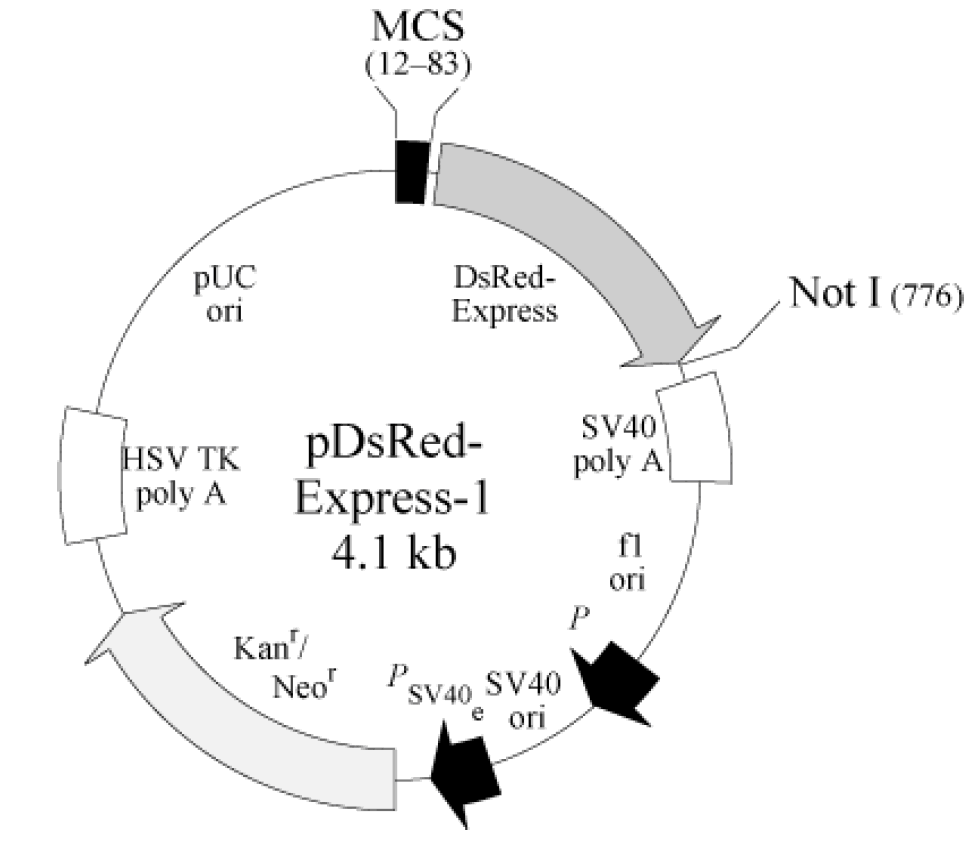
pDsRed-Express-1 is a promoterless mammalian expression vector that can be used to monitor transcription from
different promoters and promoter/enhancer combinations inserted into the multiple cloning site (MCS). It encodes DsRedExpress,
a variant of Discosoma sp. red fluorescent protein (DsRed; 1). DsRed-Express contains nine amino acid
substitutions which improve the solubility of the protein, reduce the time from transfection to detection of red
fluorescence, and decrease the level of residual green emission (2). When DsRed-Express is expressed in mammalian cell
cultures, red-emitting cells can be detected by either fluorescence microscopy or flow cytometry 8–12 hours after
transfection. Although DsRed-Express most likely forms the same tetrameric structure as wild-type DsRed, DsRedExpress
displays a reduced tendency to aggregate (2). The DsRed-Express coding sequence is human codon-optimized for
high expression in mammalian cells (3).
The sequence upstream of the DsRed-Express gene has been converted to a Kozak consensus sequence (4) to enhance
translation efficiency in eukaryotic cells. SV40 polyadenylation signals downstream of the DsRed-Express gene direct
proper processing of the 3' end of the DsRed-Express mRNA. The vector backbone contains an SV40 origin for
replication in mammalian cells expressing the SV40 T antigen, a pUC origin of replication for propagation in E. coli, and
an f1 origin for single-stranded DNA production. A neomycin-resistance cassette (Neor) allows stably transfected
eukaryotic cells to be selected using G418. This cassette consists of the SV40 early promoter, the neomycin/kanamycin
resistance gene of Tn5, and polyadenylation signals from the Herpes simplex virus thymidine kinase (HSV TK) gene. A
bacterial promoter upstream of the cassette expresses kanamycin resistance in E. coli.
Promoters should be cloned into the pDsRed-Express-1 MCS upstream from the DsRed-Express coding sequence.
Without addition of a functional promoter, this vector will not express DsRed-Express. The recombinant DsRed-Express
vector can be transfected into mammalian cells using any standard transfection method. If required, stable transfectants
can be selected using G418 (5).
LOCUS pDsRed-Express-1 4107 bp DNA SYN
DEFINITION pDsRed-Express-1
ACCESSION
KEYWORDS
SOURCE
ORGANISM other sequences; artificial sequences; vectors.
FEATURES Location/Qualifiers
source 1..4107
/organism="pDsRed-Express-1"
/mol_type="other DNA"
CDS complement(25..804)
/label="ORF frame 1"
/translation="MADYDLESRPLQEQVVAALGALVLLHDGVVLVVGGDVQLGVHVV
VAGQLHGLLGHVDGLELHQVVAAVLQLQGLVDLALQHAVAGVQALGGGLPAHSLLLHY
GAVGGEVHADELHLVDEGAVLQGGVLGHGHHAAVLEVHHALPLEALGEGQLLVVGDVG
GVLHVHLGAVLELGGQDVPGEGQGAALGHLQLGGLGALVGAALALALDLELVAVHGAL
HAHLEAHELLDDVLGGGHGGDRWIPGPRYRRLQNSKLELEI*"
gene 97..774
/label="DsRed_Express_2"
/gene="DsRed_Express_2"
/translation="MADYDLESRPLQEQVVAALGALVLLHDGVVLVVGGDVQLGVHVV
VAGQLHGLLGHVDGLELHQVVAAVLQLQGLVDLALQHAVAGVQALGGGLPAHSLLLHY
GAVGGEVHADELHLVDEGAVLQGGVLGHGHHAAVLEVHHALPLEALGEGQLLVVGDVG
GVLHVHLGAVLELGGQDVPGEGQGAALGHLQLGGLGALVGAALALALDLELVAVHGAL
HAHLEAHELLDDVLGGGHGGDRWIPGPRYRRLQNSKLELEI*"
CDS 97..774
/label="ORF frame 1"
/translation="MASSEDVIKEFMRFKVRMEGSVNGHEFEIEGEGEGRPYEGTQTA
KLKVTKGGPLPFAWDILSPQFQYGSKVYVKHPADIPDYKKLSFPEGFKWERVMNFEDG
GVVTVTQDSSLQDGSFIYKVKFIGVNFPSDGPVMQKKTMGWEASTERLYPRDGVLKGE
IHKALKLKDGGHYLVEFKSIYMAKKPVQLPGYYYVDSKLDITSHNEDYTIVEQYERAE
GRHHLFL*"
gene 100..771
/label="DsRed_Express"
/gene="DsRed_Express"
/translation="MASSEDVIKEFMRFKVRMEGSVNGHEFEIEGEGEGRPYEGTQTA
KLKVTKGGPLPFAWDILSPQFQYGSKVYVKHPADIPDYKKLSFPEGFKWERVMNFEDG
GVVTVTQDSSLQDGSFIYKVKFIGVNFPSDGPVMQKKTMGWEASTERLYPRDGVLKGE
IHKALKLKDGGHYLVEFKSIYMAKKPVQLPGYYYVDSKLDITSHNEDYTIVEQYERAE
GRHHLFL*"
misc_feature complement(277..297)
/label="dsRed1_N_primer"
/translation="MASSEDVIKEFMRFKVRMEGSVNGHEFEIEGEGEGRPYEGTQTA
KLKVTKGGPLPFAWDILSPQFQYGSKVYVKHPADIPDYKKLSFPEGFKWERVMNFEDG
GVVTVTQDSSLQDGSFIYKVKFIGVNFPSDGPVMQKKTMGWEASTERLYPRDGVLKGE
IHKALKLKDGGHYLVEFKSIYMAKKPVQLPGYYYVDSKLDITSHNEDYTIVEQYERAE
GRHHLFL*"
misc_feature 689..712
/label="dsRed1_C_primer"
/translation="MASSEDVIKEFMRFKVRMEGSVNGHEFEIEGEGEGRPYEGTQTA
KLKVTKGGPLPFAWDILSPQFQYGSKVYVKHPADIPDYKKLSFPEGFKWERVMNFEDG
GVVTVTQDSSLQDGSFIYKVKFIGVNFPSDGPVMQKKTMGWEASTERLYPRDGVLKGE
IHKALKLKDGGHYLVEFKSIYMAKKPVQLPGYYYVDSKLDITSHNEDYTIVEQYERAE
GRHHLFL*"
misc_feature 986..1005
/label="EBV_rev_primer"
/translation="MASSEDVIKEFMRFKVRMEGSVNGHEFEIEGEGEGRPYEGTQTA
KLKVTKGGPLPFAWDILSPQFQYGSKVYVKHPADIPDYKKLSFPEGFKWERVMNFEDG
GVVTVTQDSSLQDGSFIYKVKFIGVNFPSDGPVMQKKTMGWEASTERLYPRDGVLKGE
IHKALKLKDGGHYLVEFKSIYMAKKPVQLPGYYYVDSKLDITSHNEDYTIVEQYERAE
GRHHLFL*"
rep_origin complement(1155..1461)
/label="f1_origin"
/translation="MASSEDVIKEFMRFKVRMEGSVNGHEFEIEGEGEGRPYEGTQTA
KLKVTKGGPLPFAWDILSPQFQYGSKVYVKHPADIPDYKKLSFPEGFKWERVMNFEDG
GVVTVTQDSSLQDGSFIYKVKFIGVNFPSDGPVMQKKTMGWEASTERLYPRDGVLKGE
IHKALKLKDGGHYLVEFKSIYMAKKPVQLPGYYYVDSKLDITSHNEDYTIVEQYERAE
GRHHLFL*"
promoter 1540..1568
/label="AmpR_promoter"
/translation="MASSEDVIKEFMRFKVRMEGSVNGHEFEIEGEGEGRPYEGTQTA
KLKVTKGGPLPFAWDILSPQFQYGSKVYVKHPADIPDYKKLSFPEGFKWERVMNFEDG
GVVTVTQDSSLQDGSFIYKVKFIGVNFPSDGPVMQKKTMGWEASTERLYPRDGVLKGE
IHKALKLKDGGHYLVEFKSIYMAKKPVQLPGYYYVDSKLDITSHNEDYTIVEQYERAE
GRHHLFL*"
misc_feature complement(1634..1654)
/label="pBABE_3_primer"
/translation="MASSEDVIKEFMRFKVRMEGSVNGHEFEIEGEGEGRPYEGTQTA
KLKVTKGGPLPFAWDILSPQFQYGSKVYVKHPADIPDYKKLSFPEGFKWERVMNFEDG
GVVTVTQDSSLQDGSFIYKVKFIGVNFPSDGPVMQKKTMGWEASTERLYPRDGVLKGE
IHKALKLKDGGHYLVEFKSIYMAKKPVQLPGYYYVDSKLDITSHNEDYTIVEQYERAE
GRHHLFL*"
misc_feature complement(1640..1855)
/label="SV40_enhancer"
/translation="MASSEDVIKEFMRFKVRMEGSVNGHEFEIEGEGEGRPYEGTQTA
KLKVTKGGPLPFAWDILSPQFQYGSKVYVKHPADIPDYKKLSFPEGFKWERVMNFEDG
GVVTVTQDSSLQDGSFIYKVKFIGVNFPSDGPVMQKKTMGWEASTERLYPRDGVLKGE
IHKALKLKDGGHYLVEFKSIYMAKKPVQLPGYYYVDSKLDITSHNEDYTIVEQYERAE
GRHHLFL*"
promoter 1652..1920
/label="SV40_promoter"
/translation="MASSEDVIKEFMRFKVRMEGSVNGHEFEIEGEGEGRPYEGTQTA
KLKVTKGGPLPFAWDILSPQFQYGSKVYVKHPADIPDYKKLSFPEGFKWERVMNFEDG
GVVTVTQDSSLQDGSFIYKVKFIGVNFPSDGPVMQKKTMGWEASTERLYPRDGVLKGE
IHKALKLKDGGHYLVEFKSIYMAKKPVQLPGYYYVDSKLDITSHNEDYTIVEQYERAE
GRHHLFL*"
rep_origin 1819..1896
/label="SV40_origin"
/translation="MASSEDVIKEFMRFKVRMEGSVNGHEFEIEGEGEGRPYEGTQTA
KLKVTKGGPLPFAWDILSPQFQYGSKVYVKHPADIPDYKKLSFPEGFKWERVMNFEDG
GVVTVTQDSSLQDGSFIYKVKFIGVNFPSDGPVMQKKTMGWEASTERLYPRDGVLKGE
IHKALKLKDGGHYLVEFKSIYMAKKPVQLPGYYYVDSKLDITSHNEDYTIVEQYERAE
GRHHLFL*"
misc_feature 1881..1900
/label="SV40pro_F_primer"
/translation="MASSEDVIKEFMRFKVRMEGSVNGHEFEIEGEGEGRPYEGTQTA
KLKVTKGGPLPFAWDILSPQFQYGSKVYVKHPADIPDYKKLSFPEGFKWERVMNFEDG
GVVTVTQDSSLQDGSFIYKVKFIGVNFPSDGPVMQKKTMGWEASTERLYPRDGVLKGE
IHKALKLKDGGHYLVEFKSIYMAKKPVQLPGYYYVDSKLDITSHNEDYTIVEQYERAE
GRHHLFL*"
CDS 2003..2797
/label="ORF frame 2"
/translation="MIEQDGLHAGSPAAWVERLFGYDWAQQTIGCSDAAVFRLSAQGR
PVLFVKTDLSGALNELQDEAARLSWLATTGVPCAAVLDVVTEAGRDWLLLGEVPGQDL
LSSHLAPAEKVSIMADAMRRLHTLDPATCPFDHQAKHRIERARTRMEAGLVDQDDLDE
EHQGLAPAELFARLKASMPDGEDLVVTHGDACLPNIMVENGRFSGFIDCGRLGVADRY
QDIALATRDIAEELGGEWADRFLVLYGIAAPDSQRIAFYRLLDEFF*"
gene 2006..2794
/label="NeoR/KanR"
/gene="NeoR/KanR"
/translation="MIEQDGLHAGSPAAWVERLFGYDWAQQTIGCSDAAVFRLSAQGR
PVLFVKTDLSGALNELQDEAARLSWLATTGVPCAAVLDVVTEAGRDWLLLGEVPGQDL
LSSHLAPAEKVSIMADAMRRLHTLDPATCPFDHQAKHRIERARTRMEAGLVDQDDLDE
EHQGLAPAELFARLKASMPDGEDLVVTHGDACLPNIMVENGRFSGFIDCGRLGVADRY
QDIALATRDIAEELGGEWADRFLVLYGIAAPDSQRIAFYRLLDEFF*"
CDS complement(2312..2848)
/label="ORF frame 3"
/translation="MAGWASLGRSFRTPESRSEELVKKAIEGDALRIGSGDTVKHEEA
VSPFAAKLFSNITGSQRYVLIAVRHTQPATVDESRKAAIFHHDIRQAGIAMGHDEILA
VGHARLEPGEQFGWREPLMLFVQIILIDKTGFHPSTCSLDAMFRLVVEWAGSRIKRMQ
PPHCISHDGYFLGRSKVR*"
terminator 2972..3241
/label="TK_PA_terminator"
/translation="MAGWASLGRSFRTPESRSEELVKKAIEGDALRIGSGDTVKHEEA
VSPFAAKLFSNITGSQRYVLIAVRHTQPATVDESRKAAIFHHDIRQAGIAMGHDEILA
VGHARLEPGEQFGWREPLMLFVQIILIDKTGFHPSTCSLDAMFRLVVEWAGSRIKRMQ
PPHCISHDGYFLGRSKVR*"
rep_origin 3389..4008
/label="pBR322_origin"
/translation="MAGWASLGRSFRTPESRSEELVKKAIEGDALRIGSGDTVKHEEA
VSPFAAKLFSNITGSQRYVLIAVRHTQPATVDESRKAAIFHHDIRQAGIAMGHDEILA
VGHARLEPGEQFGWREPLMLFVQIILIDKTGFHPSTCSLDAMFRLVVEWAGSRIKRMQ
PPHCISHDGYFLGRSKVR*"
ORIGIN
1 TAGTTATTAC TAGCGCTACC GGACTCAGAT CTCGAGCTCA AGCTTCGAAT TCTGCAGTCG
61 ACGGTACCGC GGGCCCGGGA TCCACCGGTC GCCACCATGG CCTCCTCCGA GGACGTCATC
121 AAGGAGTTCA TGCGCTTCAA GGTGCGCATG GAGGGCTCCG TGAACGGCCA CGAGTTCGAG
181 ATCGAGGGCG AGGGCGAGGG CCGCCCCTAC GAGGGCACCC AGACCGCCAA GCTGAAGGTG
241 ACCAAGGGCG GCCCCCTGCC CTTCGCCTGG GACATCCTGT CCCCCCAGTT CCAGTACGGC
301 TCCAAGGTGT ACGTGAAGCA CCCCGCCGAC ATCCCCGACT ACAAGAAGCT GTCCTTCCCC
361 GAGGGCTTCA AGTGGGAGCG CGTGATGAAC TTCGAGGACG GCGGCGTGGT GACCGTGACC
421 CAGGACTCCT CCCTGCAGGA CGGCTCCTTC ATCTACAAGG TGAAGTTCAT CGGCGTGAAC
481 TTCCCCTCCG ACGGCCCCGT AATGCAGAAG AAGACTATGG GCTGGGAGGC CTCCACCGAG
541 CGCCTGTACC CCCGCGACGG CGTGCTGAAG GGCGAGATCC ACAAGGCCCT GAAGCTGAAG
601 GACGGCGGCC ACTACCTGGT GGAGTTCAAG TCCATCTACA TGGCCAAGAA GCCCGTGCAG
661 CTGCCCGGCT ACTACTACGT GGACTCCAAG CTGGACATCA CCTCCCACAA CGAGGACTAC
721 ACCATCGTGG AGCAGTACGA GCGCGCCGAG GGCCGCCACC ACCTGTTCCT GTAGCGGCCG
781 CGACTCTAGA TCATAATCAG CCATACCACA TTTGTAGAGG TTTTACTTGC TTTAAAAAAC
841 CTCCCACACC TCCCCCTGAA CCTGAAACAT AAAATGAATG CAATTGTTGT TGTTAACTTG
901 TTTATTGCAG CTTATAATGG TTACAAATAA AGCAATAGCA TCACAAATTT CACAAATAAA
961 GCATTTTTTT CACTGCATTC TAGTTGTGGT TTGTCCAAAC TCATCAATGT ATCTTAAGGC
1021 GTAAATTGTA AGCGTTAATA TTTTGTTAAA ATTCGCGTTA AATTTTTGTT AAATCAGCTC
1081 ATTTTTTAAC CAATAGGCCG AAATCGGCAA AATCCCTTAT AAATCAAAAG AATAGACCGA
1141 GATAGGGTTG AGTGTTGTTC CAGTTTGGAA CAAGAGTCCA CTATTAAAGA ACGTGGACTC
1201 CAACGTCAAA GGGCGAAAAA CCGTCTATCA GGGCGATGGC CCACTACGTG AACCATCACC
1261 CTAATCAAGT TTTTTGGGGT CGAGGTGCCG TAAAGCACTA AATCGGAACC CTAAAGGGAG
1321 CCCCCGATTT AGAGCTTGAC GGGGAAAGCC GGCGAACGTG GCGAGAAAGG AAGGGAAGAA
1381 AGCGAAAGGA GCGGGCGCTA GGGCGCTGGC AAGTGTAGCG GTCACGCTGC GCGTAACCAC
1441 CACACCCGCC GCGCTTAATG CGCCGCTACA GGGCGCGTCA GGTGGCACTT TTCGGGGAAA
1501 TGTGCGCGGA ACCCCTATTT GTTTATTTTT CTAAATACAT TCAAATATGT ATCCGCTCAT
1561 GAGACAATAA CCCTGATAAA TGCTTCAATA ATATTGAAAA AGGAAGAGTC CTGAGGCGGA
1621 AAGAACCAGC TGTGGAATGT GTGTCAGTTA GGGTGTGGAA AGTCCCCAGG CTCCCCAGCA
1681 GGCAGAAGTA TGCAAAGCAT GCATCTCAAT TAGTCAGCAA CCAGGTGTGG AAAGTCCCCA
1741 GGCTCCCCAG CAGGCAGAAG TATGCAAAGC ATGCATCTCA ATTAGTCAGC AACCATAGTC
1801 CCGCCCCTAA CTCCGCCCAT CCCGCCCCTA ACTCCGCCCA GTTCCGCCCA TTCTCCGCCC
1861 CATGGCTGAC TAATTTTTTT TATTTATGCA GAGGCCGAGG CCGCCTCGGC CTCTGAGCTA
1921 TTCCAGAAGT AGTGAGGAGG CTTTTTTGGA GGCCTAGGCT TTTGCAAAGA TCGATCAAGA
1981 GACAGGATGA GGATCGTTTC GCATGATTGA ACAAGATGGA TTGCACGCAG GTTCTCCGGC
2041 CGCTTGGGTG GAGAGGCTAT TCGGCTATGA CTGGGCACAA CAGACAATCG GCTGCTCTGA
2101 TGCCGCCGTG TTCCGGCTGT CAGCGCAGGG GCGCCCGGTT CTTTTTGTCA AGACCGACCT
2161 GTCCGGTGCC CTGAATGAAC TGCAAGACGA GGCAGCGCGG CTATCGTGGC TGGCCACGAC
2221 GGGCGTTCCT TGCGCAGCTG TGCTCGACGT TGTCACTGAA GCGGGAAGGG ACTGGCTGCT
2281 ATTGGGCGAA GTGCCGGGGC AGGATCTCCT GTCATCTCAC CTTGCTCCTG CCGAGAAAGT
2341 ATCCATCATG GCTGATGCAA TGCGGCGGCT GCATACGCTT GATCCGGCTA CCTGCCCATT
2401 CGACCACCAA GCGAAACATC GCATCGAGCG AGCACGTACT CGGATGGAAG CCGGTCTTGT
2461 CGATCAGGAT GATCTGGACG AAGAGCATCA GGGGCTCGCG CCAGCCGAAC TGTTCGCCAG
2521 GCTCAAGGCG AGCATGCCCG ACGGCGAGGA TCTCGTCGTG ACCCATGGCG ATGCCTGCTT
2581 GCCGAATATC ATGGTGGAAA ATGGCCGCTT TTCTGGATTC ATCGACTGTG GCCGGCTGGG
2641 TGTGGCGGAC CGCTATCAGG ACATAGCGTT GGCTACCCGT GATATTGCTG AAGAGCTTGG
2701 CGGCGAATGG GCTGACCGCT TCCTCGTGCT TTACGGTATC GCCGCTCCCG ATTCGCAGCG
2761 CATCGCCTTC TATCGCCTTC TTGACGAGTT CTTCTGAGCG GGACTCTGGG GTTCGAAATG
2821 ACCGACCAAG CGACGCCCAA CCTGCCATCA CGAGATTTCG ATTCCACCGC CGCCTTCTAT
2881 GAAAGGTTGG GCTTCGGAAT CGTTTTCCGG GACGCCGGCT GGATGATCCT CCAGCGCGGG
2941 GATCTCATGC TGGAGTTCTT CGCCCACCCT AGGGGGAGGC TAACTGAAAC ACGGAAGGAG
3001 ACAATACCGG AAGGAACCCG CGCTATGACG GCAATAAAAA GACAGAATAA AACGCACGGT
3061 GTTGGGTCGT TTGTTCATAA ACGCGGGGTT CGGTCCCAGG GCTGGCACTC TGTCGATACC
3121 CCACCGAGAC CCCATTGGGG CCAATACGCC CGCGTTTCTT CCTTTTCCCC ACCCCACCCC
3181 CCAAGTTCGG GTGAAGGCCC AGGGCTCGCA GCCAACGTCG GGGCGGCAGG CCCTGCCATA
3241 GCCTCAGGTT ACTCATATAT ACTTTAGATT GATTTAAAAC TTCATTTTTA ATTTAAAAGG
3301 ATCTAGGTGA AGATCCTTTT TGATAATCTC ATGACCAAAA TCCCTTAACG TGAGTTTTCG
3361 TTCCACTGAG CGTCAGACCC CGTAGAAAAG ATCAAAGGAT CTTCTTGAGA TCCTTTTTTT
3421 CTGCGCGTAA TCTGCTGCTT GCAAACAAAA AAACCACCGC TACCAGCGGT GGTTTGTTTG
3481 CCGGATCAAG AGCTACCAAC TCTTTTTCCG AAGGTAACTG GCTTCAGCAG AGCGCAGATA
3541 CCAAATACTG TCCTTCTAGT GTAGCCGTAG TTAGGCCACC ACTTCAAGAA CTCTGTAGCA
3601 CCGCCTACAT ACCTCGCTCT GCTAATCCTG TTACCAGTGG CTGCTGCCAG TGGCGATAAG
3661 TCGTGTCTTA CCGGGTTGGA CTCAAGACGA TAGTTACCGG ATAAGGCGCA GCGGTCGGGC
3721 TGAACGGGGG GTTCGTGCAC ACAGCCCAGC TTGGAGCGAA CGACCTACAC CGAACTGAGA
3781 TACCTACAGC GTGAGCTATG AGAAAGCGCC ACGCTTCCCG AAGGGAGAAA GGCGGACAGG
3841 TATCCGGTAA GCGGCAGGGT CGGAACAGGA GAGCGCACGA GGGAGCTTCC AGGGGGAAAC
3901 GCCTGGTATC TTTATAGTCC TGTCGGGTTT CGCCACCTCT GACTTGAGCG TCGATTTTTG
3961 TGATGCTCGT CAGGGGGGCG GAGCCTATGG AAAAACGCCA GCAACGCGGC CTTTTTACGG
4021 TTCCTGGCCT TTTGCTGGCC TTTTGCTCAC ATGTTCTTTC CTGCGTTATC CCCTGATTCT
4081 GTGGATAACC GTATTACCGC CATGCAT
//